SETD1B-mediated broad H3K4me3 controls proper temporal patterns of gene expression critical for spermatid development
IF 25.9
1区 生物学
Q1 CELL BIOLOGY
引用次数: 0
Abstract
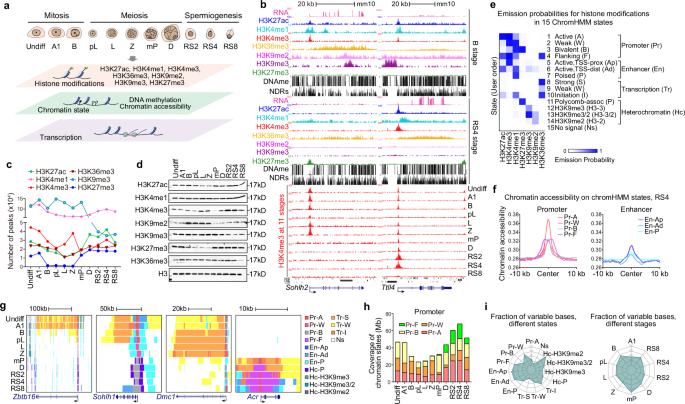
Epigenetic programming governs cell fate determination during development through intricately controlling sequential gene activation and repression. Although H3K4me3 is widely recognized as a hallmark of gene activation, its role in modulating transcription output and timing within a continuously developing system remains poorly understood. In this study, we provide a detailed characterization of the epigenomic landscapes in developing male germ cells. We identified thousands of spermatid-specific broad H3K4me3 domains regulated by the SETD1B-RFX2 axis, representing a previously underappreciated form of H3K4me3. These domains, overlapping with H3K27ac-marked enhancers and promoters, play critical roles in orchestrating robust transcription and accurate temporal control of gene expression. Mechanistically, these broad H3K4me3 compete effectively with regular H3K4me3 for transcriptional machinery, thereby ensuring robust levels and precise timing of master gene expression in mouse spermiogenesis. Disruption of this mechanism compromises the accuracy of transcription dosage and timing, ultimately impairing spermiogenesis. Additionally, we unveil remarkable changes in the distribution of heterochromatin marks, including H3K27me3 and H3K9me2, during the mitosis-to-meiosis transition and completion of meiotic recombination, which closely correlates with gene silencing. This work underscores the highly orchestrated epigenetic regulation in spermatogenesis, highlighting the previously unrecognized role of Setd1b in the formation of broad H3K4me3 domains and transcriptional control, and provides an invaluable resource for future studies toward the elucidation of spermatogenesis.

setd1b介导的广泛H3K4me3控制对精子发育至关重要的基因表达的适当时间模式
表观遗传编程通过复杂地控制序列基因激活和抑制来决定细胞在发育过程中的命运。尽管H3K4me3被广泛认为是基因激活的标志,但它在持续发育的系统中调节转录输出和时间的作用仍然知之甚少。在这项研究中,我们提供了发育中的男性生殖细胞的表观基因组景观的详细表征。我们发现了数千个由SETD1B-RFX2轴调节的精细胞特异性宽H3K4me3结构域,代表了以前未被重视的H3K4me3形式。这些结构域与h3k27ac标记的增强子和启动子重叠,在协调稳健的转录和精确的基因表达时间控制中发挥关键作用。从机制上讲,这些广泛的H3K4me3与常规的H3K4me3有效地竞争转录机制,从而确保小鼠精子发生中主基因表达的稳定水平和精确时间。这种机制的破坏会损害转录剂量和时间的准确性,最终损害精子发生。此外,我们揭示了异染色质标记H3K27me3和H3K9me2在有丝分裂到减数分裂过渡和减数分裂重组完成过程中的显著变化,这与基因沉默密切相关。这项工作强调了精子发生过程中高度协调的表观遗传调控,强调了Setd1b在广泛H3K4me3结构域的形成和转录控制中的作用,并为未来研究精子发生提供了宝贵的资源。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Cell Research
生物-细胞生物学
CiteScore
53.90
自引率
0.70%
发文量
2420
审稿时长
2.3 months
期刊介绍:
Cell Research (CR) is an international journal published by Springer Nature in partnership with the Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences (CAS). It focuses on publishing original research articles and reviews in various areas of life sciences, particularly those related to molecular and cell biology. The journal covers a broad range of topics including cell growth, differentiation, and apoptosis; signal transduction; stem cell biology and development; chromatin, epigenetics, and transcription; RNA biology; structural and molecular biology; cancer biology and metabolism; immunity and molecular pathogenesis; molecular and cellular neuroscience; plant molecular and cell biology; and omics, system biology, and synthetic biology. CR is recognized as China's best international journal in life sciences and is part of Springer Nature's prestigious family of Molecular Cell Biology journals.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


