The genomic landscape of gene-level structural variations in Japanese and global soybean Glycine max cultivars
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
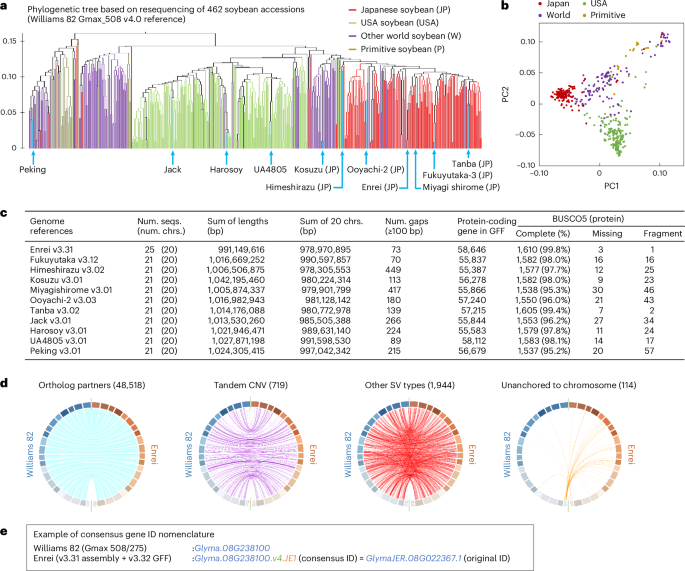
Japanese soybeans are traditionally bred to produce soy foods such as tofu, miso and boiled soybeans. Here, to investigate their distinctive genomic features, including genomic structural variations (SVs), we constructed 11 nanopore-based genome references for Japanese and other soybean lines. Our assembly-based comparative method, designated ‘Asm2sv’, identified gene-level SVs comprehensively, enabling pangenome analysis of 462 worldwide cultivars and varieties. Based on these, we identified selective sweeps between Japanese and US soybeans, one of which was the pod-shattering resistance gene PDH1. Genome-wide association studies further identified several quantitative trait loci that accounted for large-seed phenotypes of Japanese soybean lines, some of which were also close to regions of the selective sweeps, including PDH1. Notably, specific combinations of alleles, including SVs, were found to increase the seed size of some Japanese landraces. In addition to the differences in cultivation environments, distinct food processing usages might result in changes in Japanese soybean genomes. Long-read genome assemblies for seven Japanese, three North American and one primitive Glycine max cultivars highlight gene-level structural variation underlying distinct seed morphology phenotypes.

日本和全球大豆品种基因水平结构变异的基因组图谱
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


