Integrated analysis of the complete sequence of a macaque genome
IF 48.5
1区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 0
Abstract
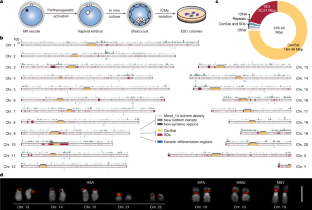
The crab-eating macaques (Macaca fascicularis) and rhesus macaques (Macaca mulatta) are pivotal in biomedical and evolutionary research1–3. However, their genomic complexity and interspecies genetic differences remain unclear4. Here, we present a complete genome assembly of a crab-eating macaque, revealing 46% fewer segmental duplications and 3.83 times longer centromeres than those of humans5,6. We also characterize 93 large-scale genomic differences between macaques and humans at a single-base-pair resolution, highlighting their impact on gene regulation in primate evolution. Using ten long-read macaque genomes, hundreds of short-read macaque genomes and full-length transcriptome data, we identified roughly 2 Mbp of fixed-genetic variants, roughly 240 Mbp of complex loci, 16.76 Mbp genetic differentiation regions and 110 alternative splice events, potentially associated with various phenotypic differences between the two macaque species. In summary, the integrated genetic analysis enhances understanding of lineage-specific phenotypes, adaptation and primate evolution, thereby improving their biomedical applications in human disease research. A complete genome assembly of a crab-eating macaque, revealing 46% fewer segmental duplications and 3.83 times longer centromeres than those of humans, is presented, enhancing understanding of lineage-specific phenotypes, adaptation and primate evolution.

猕猴基因组完整序列的综合分析
食蟹猕猴(Macaca fascicularis)和恒河猴(Macaca mulatta)在生物医学和进化研究中起着关键作用。然而,它们的基因组复杂性和种间遗传差异仍不清楚。在这里,我们展示了一种食蟹猕猴的完整基因组组装,揭示了比人类少46%的片段重复和3.83倍长的着丝粒5,6。我们还在单碱基对分辨率下描述了猕猴和人类之间的93种大规模基因组差异,强调了它们对灵长类动物进化中基因调控的影响。利用10个长读猕猴基因组,数百个短读猕猴基因组和全长转录组数据,我们确定了大约2 Mbp的固定遗传变异,大约240 Mbp的复杂位点,16.76 Mbp的遗传分化区域和110个可选剪接事件,这些可能与两种猕猴之间的各种表型差异有关。总之,综合遗传分析增强了对谱系特异性表型、适应和灵长类动物进化的理解,从而提高了它们在人类疾病研究中的生物医学应用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature
综合性期刊-综合性期刊
CiteScore
90.00
自引率
1.20%
发文量
3652
审稿时长
3 months
期刊介绍:
Nature is a prestigious international journal that publishes peer-reviewed research in various scientific and technological fields. The selection of articles is based on criteria such as originality, importance, interdisciplinary relevance, timeliness, accessibility, elegance, and surprising conclusions. In addition to showcasing significant scientific advances, Nature delivers rapid, authoritative, insightful news, and interpretation of current and upcoming trends impacting science, scientists, and the broader public. The journal serves a dual purpose: firstly, to promptly share noteworthy scientific advances and foster discussions among scientists, and secondly, to ensure the swift dissemination of scientific results globally, emphasizing their significance for knowledge, culture, and daily life.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


