CRISPRi-ART enables functional genomics of diverse bacteriophages using RNA-binding dCas13d
IF 20.5
1区 生物学
Q1 MICROBIOLOGY
引用次数: 0
Abstract
Bacteriophages constitute one of the largest reservoirs of genes of unknown function in the biosphere. Even in well-characterized phages, the functions of most genes remain unknown. Experimental approaches to study phage gene fitness and function at genome scale are lacking, partly because phages subvert many modern functional genomics tools. Here we leverage RNA-targeting dCas13d to selectively interfere with protein translation and to measure phage gene fitness at a transcriptome-wide scale. We find CRISPR Interference through Antisense RNA-Targeting (CRISPRi-ART) to be effective across phage phylogeny, from model ssRNA, ssDNA and dsDNA phages to nucleus-forming jumbo phages. Using CRISPRi-ART, we determine a conserved role of diverse rII homologues in subverting phage Lambda RexAB-mediated immunity to superinfection and identify genes critical for phage fitness. CRISPRi-ART establishes a broad-spectrum phage functional genomics platform, revealing more than 90 previously unknown genes important for phage fitness. Leveraging RNA-targeting dCas13d enables selective interference with phage protein translation and facilitates measurement of phage gene fitness at a transcriptome-wide scale.

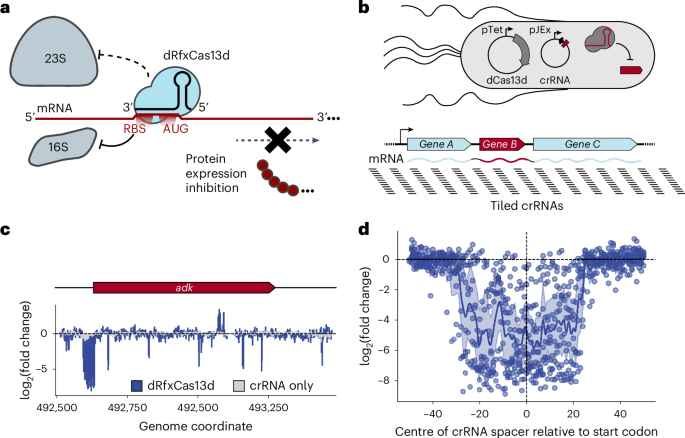
crispr - art利用rna结合dCas13d实现多种噬菌体的功能基因组学
噬菌体是生物圈中功能未知的基因的最大储存库之一。即使在特征明确的噬菌体中,大多数基因的功能仍然未知。在基因组尺度上研究噬菌体基因适应性和功能的实验方法缺乏,部分原因是噬菌体颠覆了许多现代功能基因组学工具。在这里,我们利用rna靶向dCas13d选择性地干扰蛋白质翻译,并在转录组范围内测量噬菌体基因适应度。我们发现通过反义rna靶向(CRISPR - art)进行CRISPR干扰在整个噬菌体系统发育中都是有效的,从模型ssRNA、ssDNA和dsDNA噬菌体到成核巨型噬菌体。利用crispr - art,我们确定了多种rII同源物在破坏噬菌体Lambda rexab介导的重复感染免疫中的保守作用,并鉴定了对噬菌体适应性至关重要的基因。crispr - art建立了一个广谱噬菌体功能基因组学平台,揭示了90多个以前未知的对噬菌体适应性重要的基因。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Microbiology
Immunology and Microbiology-Microbiology
CiteScore
44.40
自引率
1.10%
发文量
226
期刊介绍:
Nature Microbiology aims to cover a comprehensive range of topics related to microorganisms. This includes:
Evolution: The journal is interested in exploring the evolutionary aspects of microorganisms. This may include research on their genetic diversity, adaptation, and speciation over time.
Physiology and cell biology: Nature Microbiology seeks to understand the functions and characteristics of microorganisms at the cellular and physiological levels. This may involve studying their metabolism, growth patterns, and cellular processes.
Interactions: The journal focuses on the interactions microorganisms have with each other, as well as their interactions with hosts or the environment. This encompasses investigations into microbial communities, symbiotic relationships, and microbial responses to different environments.
Societal significance: Nature Microbiology recognizes the societal impact of microorganisms and welcomes studies that explore their practical applications. This may include research on microbial diseases, biotechnology, or environmental remediation.
In summary, Nature Microbiology is interested in research related to the evolution, physiology and cell biology of microorganisms, their interactions, and their societal relevance.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


