Developing splice-switching oligonucleotides for urea cycle disorder using an integrated diagnostic and therapeutic platform
IF 26.8
1区 医学
Q1 GASTROENTEROLOGY & HEPATOLOGY
引用次数: 0
Abstract
Backgrounds & Aims
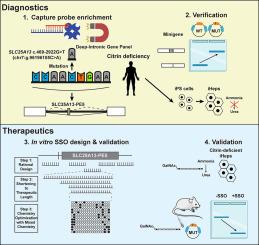
Citrin deficiency (CD) is an autosomal recessive urea cycle disorder caused by biallelic loss-of-function variants in the SLC25A13 gene, leading to life-threatening hyperammonemia and hypoglycemia. Variants in deep introns can cause genetic diseases by altering splicing and are often missed by current diagnostic tools. Splice-switching oligonucleotides (SSOs) can resolve certain intronic variants, but patients harboring such variants need to be identified. We present a lean workflow from molecular diagnostics to SSO development to resolve splice-altering variants in deep introns that is applicable to other genetic disorders.Methods
A deep intronic gene panel was designed to identify deep intronic variants. SSOs were then developed and validated in vitro using minigene assay and induced hepatocytes, and target engagement was verified in vivo by hydrodynamic tail vein injection of minigenes and SSOs.Results
With the Deep Intronic-Gene Panel and RNA analysis, we identified a novel SLC25A13 c.469-2922G>T variant that promotes the inclusion of a premature stop codon-containing pseudo-exon, SLC25A13-PE5, thereby causing CD. By a stepwise rational SSO design approach, we identified potent candidates inhibiting SLC25A13-PE5 at EC50 <2nM in vitro. Upon conjugating the SSOs with N-acetylgalactosamine (GalNAc), they were validated to rescue normal protein expression and restore ureagenesis and ammonia clearance, key urea cycle functions, in patient-derived induced hepatocytes. In vivo on-target efficacy of the clinical GalNAc-SSO candidate, in the absence of acute toxicity and inflammation, was observed in a mouse model with exogenous hepatic minigene expression.Conclusions
Our data validates a platform to redefine molecular diagnosis of urea cycle disorders and provides a proof-of-concept for precision therapy of CD patients whose only effective treatment is liver transplantation.IMPACT & IMPLICATIONS
Deep intronic variants are common causes of genetic diseases that are commonly neglected. In this study, we demonstrate an integrated precision diagnostic and therapeutic approach for UCD. Specifically, we focus on CD, going from the discovery of a novel splice variant in the SLC25A13 gene with our novel UCD Deep Intronic-Gene Panel, to the development and in vivo validation of an efficacious SSO candidate for the pathogenic splice variant. We envision the possibility of extrapolating this pipeline to the diagnosis and development of treatments for other rare genetic diseases.
求助全文
约1分钟内获得全文
求助全文
来源期刊

Journal of Hepatology
医学-胃肠肝病学
CiteScore
46.10
自引率
4.30%
发文量
2325
审稿时长
30 days
期刊介绍:
The Journal of Hepatology is the official publication of the European Association for the Study of the Liver (EASL). It is dedicated to presenting clinical and basic research in the field of hepatology through original papers, reviews, case reports, and letters to the Editor. The Journal is published in English and may consider supplements that pass an editorial review.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


