Selective deuteration of an RNA:RNA complex for structural analysis using small-angle scattering
IF 4.3
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
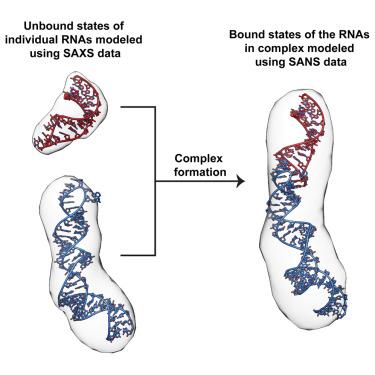
The structures of RNA:RNA complexes regulate many biological processes. Despite their importance, protein-free RNA:RNA complexes represent a tiny fraction of experimentally determined structures. Here, we describe a joint small-angle X-ray and neutron scattering (SAXS/SANS) approach to structurally interrogate conformational changes in a model RNA:RNA complex. Using SAXS, we measured the solution structures of the individual RNAs and of the overall RNA:RNA complex. With SANS, we demonstrate, as a proof of principle, that isotope labeling and contrast matching (CM) can be combined to probe the bound state structure of an RNA within a selectively deuterated RNA:RNA complex. Furthermore, we show that experimental scattering data can validate and improve predicted AlphaFold 3 RNA:RNA complex structures to reflect its solution structure. Our work demonstrates that in silico modeling, SAXS, and CM-SANS can be used in concert to directly analyze conformational changes within RNAs when in complex, enhancing our understanding of RNA structure in functional assemblies.
RNA的选择性氘化:用小角度散射进行结构分析的RNA复合体
RNA的结构:RNA复合物调节许多生物过程。尽管它们很重要,但无蛋白RNA:RNA复合物只占实验确定结构的一小部分。在这里,我们描述了一种联合小角x射线和中子散射(SAXS/SANS)方法,以结构上询问模型RNA:RNA复合物的构象变化。使用SAXS,我们测量了单个RNA和整体RNA:RNA复合物的溶液结构。通过SANS,我们证明,作为原理证明,同位素标记和对比匹配(CM)可以结合起来探测选择性氘化RNA:RNA复合物内RNA的结合态结构。此外,我们表明实验散射数据可以验证和改进预测的AlphaFold 3 RNA:RNA复合物结构,以反映其溶液结构。我们的工作表明,在硅模型中,SAXS和CM-SANS可以协同使用,直接分析RNA复杂时的构象变化,增强我们对功能组装中RNA结构的理解。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


