CryoVIA: An image analysis toolkit for the quantification of membrane structures from cryo-EM micrographs
IF 4.3
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
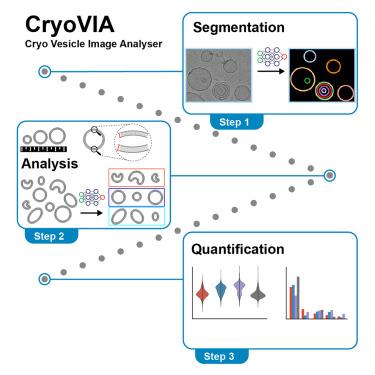
Imaging of lipid structures and associated protein complexes using cryoelectron microscopy (cryo-EM) is a common visualization and structure determination technique. The quantitative analysis of the membrane structures, however, is not routine and time consuming in particular when large amounts of data are involved. Here, we introduce the automated image-processing software cryo-vesicle image analyzer (CryoVIA) that parametrizes lipid structures of large datasets from cryo-EM images. This toolkit combines segmentation, structure identification with methods to automatically perform a large-scale data analysis of local and global membrane properties such as bilayer thickness, size, and curvature including membrane shape classifications. We included analyses of exemplary datasets of different lipid compositions and protein-induced lipid changes through an endosomal sorting complexes required for transport III (ESCRT-III) membrane remodeling protein. The toolkit opens new possibilities to systematically study structural properties of membrane structures and their modifications from cryo-EM images.
CryoVIA:一个图像分析工具包,用于从冷冻电镜显微照片中定量膜结构
使用低温电子显微镜(cryo-EM)成像脂质结构和相关蛋白复合物是一种常见的可视化和结构测定技术。膜结构的定量分析,然而,不是常规的和耗时的,特别是当涉及大量的数据。在这里,我们介绍了自动图像处理软件冷冻囊泡图像分析仪(CryoVIA),它可以从冷冻电镜图像中参数化大数据集的脂质结构。该工具包结合了分割、结构识别和自动执行局部和全局膜属性(如双层厚度、大小和曲率)的大规模数据分析方法,包括膜形状分类。我们分析了不同脂质组成的典型数据集,以及通过转运III (ESCRT-III)膜重塑蛋白所需的内体分选复合物引起的蛋白质脂质变化。该工具包为系统地研究膜结构的结构特性及其从低温电镜图像中的修饰开辟了新的可能性。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


