Transposon proliferation drives genome architecture and regulatory evolution in wild and domesticated peppers
IF 15.8
1区 生物学
Q1 PLANT SCIENCES
引用次数: 0
Abstract
Pepper (Capsicum spp.) is a widely consumed vegetable with exceptionally large genomes in Solanaceae, yet its genomic evolutionary history remains largely unknown. Here we present 11 high-quality Capsicum genome assemblies, including two gap-free genomes, covering four wild and all five domesticated pepper species. We reconstructed the ancestral karyotype and inferred the evolutionary trajectory of peppers. The expanded and variable genome sizes were attributed to differential transposable element accumulations, which shaped 3D chromatin architecture and introduced mutations associated with traits such as fruit orientation and colour. Using a chromatin accessibility atlas of Capsicum, we highlight the influence of transposable elements on regulatory element evolution. Furthermore, by constructing a haploblock map of 124 pepper core germplasms, we uncover frequent introgressions that facilitate the formation of sweet blocky pepper and the acquisition of important traits such as resistance to pepper mild mottle virus. These findings on the genomic and functional evolution of Capsicum will benefit pepper breeding. This study presents 11 telomere-to-telomere genomes of wild and domesticated pepper, highlights how transposable elements have shaped the evolution of genome structure and regulatory elements and identifies structural variations and introgressions associated with key traits in cultivated pepper.

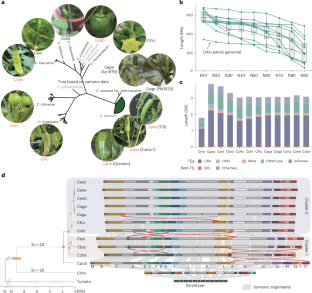
转座子增殖驱动野生和驯化辣椒的基因组结构和调控进化
辣椒(Capsicum spp.)是一种广泛食用的蔬菜,在茄科中具有特别大的基因组,但其基因组进化史在很大程度上仍然未知。在这里,我们展示了11个高质量的辣椒基因组组合,包括两个无缺口基因组,涵盖了4个野生辣椒和所有5个驯化辣椒物种。我们重建了祖先的核型,并推断了辣椒的进化轨迹。扩大和可变的基因组大小归因于不同的转座因子积累,这形成了三维染色质结构,并引入了与果实取向和颜色等性状相关的突变。利用辣椒的染色质可及性图谱,我们强调了转座元件对调控元件进化的影响。此外,通过构建124份辣椒核心种质的单倍块图谱,我们发现了促进甜块状辣椒形成和获得抗辣椒轻度斑纹病毒等重要性状的频繁渗入。这些关于辣椒基因组和功能进化的研究成果将为辣椒育种提供参考。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature Plants
PLANT SCIENCES-
CiteScore
25.30
自引率
2.20%
发文量
196
期刊介绍:
Nature Plants is an online-only, monthly journal publishing the best research on plants — from their evolution, development, metabolism and environmental interactions to their societal significance.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


