Structural and functional analysis of SAM-dependent N-methyltransferases involved in ovoselenol and ovothiol biosynthesis
IF 4.4
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
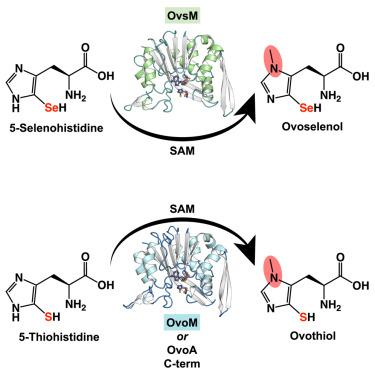
Thio/selenoimidazole Nπ-methyltransferases are an emerging family of enzymes catalyzing the final step in the production of the S/Se-containing histidine-derived antioxidants ovothiol and ovoselenol. These enzymes, prevalent in prokaryotes, show minimal sequence similarity to other methyltransferases, and the structural determinants of their reactivities remain poorly understood. Herein, we report ligand-bound crystal structures of OvsM from the ovoselenol pathway as well as a member of a previously unknown clade of standalone ovothiol-biosynthetic Nπ-methyltransferases, which we have designated OvoM. Unlike previously reported ovothiol methyltransferases, which are fused as a C-terminal domain to the sulfoxide synthase OvoA, OvoMs function independently. Comparative structural analyses reveal conserved, ligand-induced conformational changes, suggesting similar behavior in dual-domain OvoA enzymes. Mutagenesis supports a model where OvoA domain rearrangement facilitates substrate recognition via a critical Tyr residue in the domain linker. Biochemical studies identify an essential active-site Asp, likely serving as a catalytic base in the SN2-like nucleophilic substitution reaction.
求助全文
约1分钟内获得全文
求助全文
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


