Metagenomic and enzymatic mechanisms underpinning efficient water treatment of 2-ethylhexyl diphenyl phosphate (EHDPP) by the microbial consortium 8-ZY
IF 11.4
1区 环境科学与生态学
Q1 ENGINEERING, ENVIRONMENTAL
引用次数: 0
Abstract
The ubiquitous presence, potential toxicity, and persistence of 2-ethylhexyl diphenyl phosphate (EHDPP) in the environment have raised significant concerns. In this study, we successfully isolate a novel microbial consortium, named 8-ZY, and we demonstrate its remarkable ability to degrade EHDPP using an extremely low concentration of the inoculate. A total of 11 degradation metabolites were identified, including hydrolysis, hydroxylated, methylated, glucuronide-conjugated, and previously unreported byproducts, enabling us to propose new transformation pathways. Further, we unveiled the active members of the microbial consortium 8-ZY during the degradation of EHDPP. We observed the presence of diverse active populations, which included Bradyrhizobium, Rhodopseudomonas, Sphingomonas, Hyphomicrobium, Chitinophaga, Aminobacter, and Ralstonia. A metagenomic analysis revealed the presence of genes that encode phosphatase, phosphodiesterase, cytochrome P450, and hydroxylase enzymes, thus indicating their crucial role in EHDPP degradation. Furthermore, we successfully isolated Burkholderia cepacia ZY1, Sphingopyxis terrae ZY2, and Amycolatopsis ZY3 from the 8-ZY consortium, confirming their significance in EHDPP degradation and metabolite formation. These findings underscored the diversity of strains and functional genes responsible for the transformation of EHDPP within the consortium 8-ZY, highlighting the essential role of synergistic interactions during EHDPP biodegradation processes. Molecular docking and dynamics simulation suggested that alkaline phosphatase, cytochrome P450, and hydroxylase stably bonded to EHDPP within their respective active pockets, targeting distinct sites on the EHDPP molecule. These findings provide a comprehensive understanding of the transformation mechanisms of OPEs and contribute valuable insights into their fate in the environment.

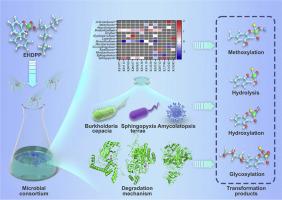
微生物联盟8-ZY对2-乙基己基磷酸二苯酯(EHDPP)进行有效水处理的宏基因组学和酶学机制
2-乙基己基二苯基磷酸(EHDPP)在环境中的普遍存在、潜在毒性和持久性引起了人们的极大关注。在这项研究中,我们成功地分离了一个新的微生物联合体,命名为8-ZY,我们证明了它使用极低浓度的接种物降解EHDPP的显著能力。共鉴定出11种降解代谢物,包括水解、羟基化、甲基化、葡萄糖醛酸缀合和以前未报道的副产物,使我们能够提出新的转化途径。此外,我们揭示了微生物联盟8-ZY在EHDPP降解过程中的活跃成员。我们观察到多种活跃种群的存在,包括慢根瘤菌、红假单胞菌、鞘单胞菌、菌丝微生物、几丁菌、氨基杆菌和拉尔斯顿菌。宏基因组分析显示存在编码磷酸酶、磷酸二酯酶、细胞色素P450和羟化酶的基因,从而表明它们在EHDPP降解中起着至关重要的作用。此外,我们成功地从8-ZY联盟中分离出洋葱伯克霍尔德菌ZY1、地鞘菌ZY2和Amycolatopsis ZY3,证实了它们在EHDPP降解和代谢物形成中的重要意义。这些发现强调了8-ZY联盟中负责EHDPP转化的菌株和功能基因的多样性,突出了协同相互作用在EHDPP生物降解过程中的重要作用。分子对接和动力学模拟表明,碱性磷酸酶、细胞色素P450和羟化酶在各自的活性口袋内稳定地与EHDPP结合,靶向EHDPP分子上的不同位点。这些发现为OPEs的转化机制提供了全面的理解,并为它们在环境中的命运提供了有价值的见解。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Water Research
环境科学-工程:环境
CiteScore
20.80
自引率
9.40%
发文量
1307
审稿时长
38 days
期刊介绍:
Water Research, along with its open access companion journal Water Research X, serves as a platform for publishing original research papers covering various aspects of the science and technology related to the anthropogenic water cycle, water quality, and its management worldwide. The audience targeted by the journal comprises biologists, chemical engineers, chemists, civil engineers, environmental engineers, limnologists, and microbiologists. The scope of the journal include:
•Treatment processes for water and wastewaters (municipal, agricultural, industrial, and on-site treatment), including resource recovery and residuals management;
•Urban hydrology including sewer systems, stormwater management, and green infrastructure;
•Drinking water treatment and distribution;
•Potable and non-potable water reuse;
•Sanitation, public health, and risk assessment;
•Anaerobic digestion, solid and hazardous waste management, including source characterization and the effects and control of leachates and gaseous emissions;
•Contaminants (chemical, microbial, anthropogenic particles such as nanoparticles or microplastics) and related water quality sensing, monitoring, fate, and assessment;
•Anthropogenic impacts on inland, tidal, coastal and urban waters, focusing on surface and ground waters, and point and non-point sources of pollution;
•Environmental restoration, linked to surface water, groundwater and groundwater remediation;
•Analysis of the interfaces between sediments and water, and between water and atmosphere, focusing specifically on anthropogenic impacts;
•Mathematical modelling, systems analysis, machine learning, and beneficial use of big data related to the anthropogenic water cycle;
•Socio-economic, policy, and regulations studies.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


