Metagenome-informed metaproteomics of the human gut microbiome, host, and dietary exposome uncovers signatures of health and inflammatory bowel disease
IF 42.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
Host-microbiome-dietary interactions play crucial roles in regulating human health, yet their direct functional assessment remains challenging. We adopted metagenome-informed metaproteomics (MIM), in mice and humans, to non-invasively explore species-level microbiome-host interactions during commensal and pathogen colonization, nutritional modification, and antibiotic-induced perturbation. Simultaneously, fecal MIM accurately characterized the nutritional exposure landscape in multiple clinical and dietary contexts. Implementation of MIM in murine auto-inflammation and in human inflammatory bowel disease (IBD) characterized a “compositional dysbiosis” and a concomitant species-specific “functional dysbiosis” driven by suppressed commensal responses to inflammatory host signals. Microbiome transfers unraveled early-onset kinetics of these host-commensal cross-responsive patterns, while predictive analyses identified candidate fecal host-microbiome IBD biomarker protein pairs outperforming S100A8/S100A9 (calprotectin). Importantly, a simultaneous fecal nutritional MIM assessment enabled the determination of IBD-related consumption patterns, dietary treatment compliance, and small intestinal digestive aberrations. Collectively, a parallelized dietary-bacterial-host MIM assessment functionally uncovers trans-kingdom interactomes shaping gastrointestinal ecology while offering personalized diagnostic and therapeutic insights into microbiome-associated disease.
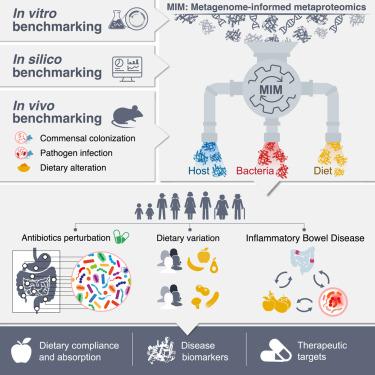
人类肠道微生物组、宿主和饮食暴露的宏基因组信息揭示了健康和炎症性肠病的特征
宿主-微生物组-饮食相互作用在调节人类健康中起着至关重要的作用,但其直接功能评估仍然具有挑战性。我们在小鼠和人类中采用了宏基因组信息宏蛋白质组学(MIM),以非侵入性地探索在共生和病原体定植、营养修饰和抗生素诱导的扰动过程中物种水平的微生物组-宿主相互作用。同时,粪便MIM准确地描述了多种临床和饮食背景下的营养暴露情况。在小鼠自身炎症和人类炎症性肠病(IBD)中,MIM的实施表现为“组成失调”和伴随的物种特异性“功能失调”,这是由对炎症宿主信号的抑制共生反应驱动的。微生物组转移揭示了这些宿主-共生交叉反应模式的早发动力学,而预测分析确定了候选粪便宿主-微生物组IBD生物标志物蛋白对,其表现优于S100A8/S100A9(钙保护蛋白)。重要的是,同时进行的粪便营养MIM评估可以确定ibd相关的消费模式、饮食治疗依从性和小肠消化异常。总的来说,一项平行的饮食-细菌-宿主MIM评估从功能上揭示了塑造胃肠道生态的跨界相互作用组,同时为微生物组相关疾病提供个性化的诊断和治疗见解。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Cell
生物-生化与分子生物学
CiteScore
110.00
自引率
0.80%
发文量
396
审稿时长
2 months
期刊介绍:
Cells is an international, peer-reviewed, open access journal that focuses on cell biology, molecular biology, and biophysics. It is affiliated with several societies, including the Spanish Society for Biochemistry and Molecular Biology (SEBBM), Nordic Autophagy Society (NAS), Spanish Society of Hematology and Hemotherapy (SEHH), and Society for Regenerative Medicine (Russian Federation) (RPO).
The journal publishes research findings of significant importance in various areas of experimental biology, such as cell biology, molecular biology, neuroscience, immunology, virology, microbiology, cancer, human genetics, systems biology, signaling, and disease mechanisms and therapeutics. The primary criterion for considering papers is whether the results contribute to significant conceptual advances or raise thought-provoking questions and hypotheses related to interesting and important biological inquiries.
In addition to primary research articles presented in four formats, Cells also features review and opinion articles in its "leading edge" section, discussing recent research advancements and topics of interest to its wide readership.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


