A widespread and ancient bacterial machinery assembles cytochrome OmcS nanowires essential for extracellular electron transfer
IF 7.2
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
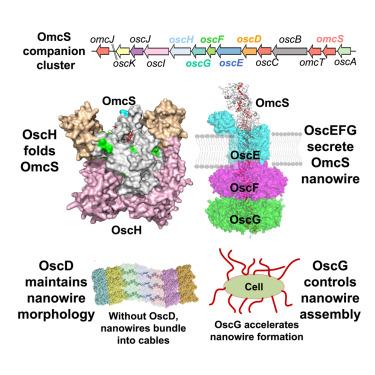
Microbial extracellular electron transfer (EET) drives various globally important environmental phenomena and has biotechnology applications. Diverse prokaryotes have been proposed to perform EET via surface-displayed “nanowires” composed of multi-heme cytochromes. However, the mechanism that enables only a few cytochromes to polymerize into nanowires is unclear. Here, we identify a highly conserved omcS-companion (osc) cluster that drives the formation of cytochrome OmcS nanowires in Geobacter sulfurreducens. Through a combination of genetic, biochemical, and biophysical methods, we establish that prolyl isomerase-containing chaperon OscH, channel-like OscEFG, and β-propeller-like OscD are involved in the folding, secretion, and morphology maintenance of OmcS nanowires, respectively. OscH and OscG can interact with OmcS. Furthermore, overexpression of oscG accelerates EET by overproducing nanowires in an ATP-dependent manner. Heme loading splits OscD; ΔoscD accelerates cell growth, bundles nanowires into cables. Our findings establish the mechanism and prevalence of a specialized and modular assembly system for nanowires across phylogenetically diverse species and environments

一种广泛而古老的细菌机器组装细胞色素OmcS纳米线,对细胞外电子转移至关重要
微生物胞外电子转移(EET)驱动着各种全球重要的环境现象,并具有生物技术应用。不同的原核生物已经被提出通过由多血红素细胞色素组成的表面显示的“纳米线”来执行EET。然而,使少数细胞色素聚合成纳米线的机制尚不清楚。在这里,我们确定了一个高度保守的OmcS -伴侣(osc)簇,该簇驱动硫还原地杆菌中细胞色素OmcS纳米线的形成。通过遗传学、生物化学和生物物理方法的结合,我们确定含有脯氨酰异构酶的伴侣蛋白OscH、通道状OscEFG和β-螺旋桨状OscD分别参与了OmcS纳米线的折叠、分泌和形态维持。OscH和OscG可以与omc交互。此外,oscG的过表达通过以atp依赖的方式过量产生纳米线来加速EET。血红素加载分裂OscD;ΔoscD加速细胞生长,将纳米线束成电缆。我们的研究结果建立了一种专门的、模块化的纳米线组装系统的机制和普遍性,这种系统可以跨越不同的物种和环境
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Cell Chemical Biology
Biochemistry, Genetics and Molecular Biology-Molecular Medicine
CiteScore
14.70
自引率
2.30%
发文量
143
期刊介绍:
Cell Chemical Biology, a Cell Press journal established in 1994 as Chemistry & Biology, focuses on publishing crucial advances in chemical biology research with broad appeal to our diverse community, spanning basic scientists to clinicians. Pioneering investigations at the chemistry-biology interface, the journal fosters collaboration between these disciplines. We encourage submissions providing significant conceptual advancements of broad interest across chemical, biological, clinical, and related fields. Particularly sought are articles utilizing chemical tools to perturb, visualize, and measure biological systems, offering unique insights into molecular mechanisms, disease biology, and therapeutics.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


