Deciphering the Coupling State-Dependent Transcription Termination in the Escherichia coli Galactose Operon
IF 2.6
2区 生物学
Q3 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
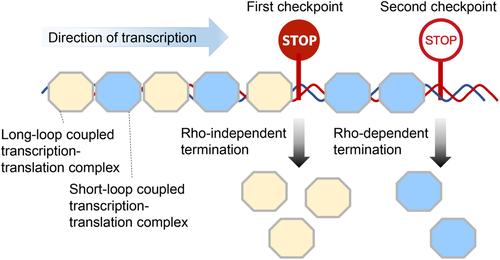
The distance between the ribosome and the RNA polymerase active centers, known as the mRNA loop length, is crucial for transcription-translation coupling. Despite the existence of multiple expressomes with varying mRNA loop lengths, their in vivo roles remain largely unexplored. This study examines the mechanisms governing transcription termination in the Escherichia coli galactose operon, revealing a crucial role in the transcription and translation coupling state. The operon utilizes both Rho-independent and Rho-dependent terminators. Our findings demonstrate that long-loop coupled transcription-translation complexes preferentially terminate at the upstream Rho-independent terminator, while short-loop complexes bypass it, terminating at the downstream Rho-dependent terminator. The efficiency of the Rho-independent terminator is enhanced by an extended U-track, suggesting a novel mechanism to overcome ribosome inhibition. These results uncover a new regulatory layer in transcription termination, challenging the traditional view of this process as random and highlighting a predetermined mechanism based on the coupling state. We propose that tandem terminators may function as regulatory checkpoints under fluctuating ribosome-RNAP coupling conditions, which can occur due to specific cellular states or factors affecting ribosome or RNAP binding efficiency. This suggests a previously overlooked mechanism that could refine transcription termination choices and expand our understanding of transcription regulation.
破译大肠杆菌半乳糖操纵子偶联状态依赖性转录终止
核糖体和RNA聚合酶活性中心之间的距离,即mRNA环长度,对转录-翻译偶联至关重要。尽管存在多种mRNA环长度不同的表达体,但它们在体内的作用仍未得到充分研究。本研究探讨了大肠杆菌半乳糖操纵子的转录终止机制,揭示了其在转录和翻译耦合状态中的关键作用。操纵子同时使用rho独立终止子和rho依赖终止子。我们的研究结果表明,长环偶联转录-翻译复合物优先终止于上游rho非依赖性终止子,而短环复合物绕过它,终止于下游rho依赖性终止子。rho非依赖性终止子的效率通过延长的u型轨道得到提高,这提示了克服核糖体抑制的新机制。这些结果揭示了转录终止中的一个新的调控层,挑战了这一过程是随机的传统观点,并强调了基于耦合状态的预定机制。我们提出串联终止子可能在波动的核糖体-RNAP耦合条件下发挥调控检查点的作用,这可能是由于特定的细胞状态或影响核糖体或RNAP结合效率的因素而发生的。这提示了一种以前被忽视的机制,可以改进转录终止选择并扩展我们对转录调控的理解。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Molecular Microbiology
生物-生化与分子生物学
CiteScore
7.20
自引率
5.60%
发文量
132
审稿时长
1.7 months
期刊介绍:
Molecular Microbiology, the leading primary journal in the microbial sciences, publishes molecular studies of Bacteria, Archaea, eukaryotic microorganisms, and their viruses.
Research papers should lead to a deeper understanding of the molecular principles underlying basic physiological processes or mechanisms. Appropriate topics include gene expression and regulation, pathogenicity and virulence, physiology and metabolism, synthesis of macromolecules (proteins, nucleic acids, lipids, polysaccharides, etc), cell biology and subcellular organization, membrane biogenesis and function, traffic and transport, cell-cell communication and signalling pathways, evolution and gene transfer. Articles focused on host responses (cellular or immunological) to pathogens or on microbial ecology should be directed to our sister journals Cellular Microbiology and Environmental Microbiology, respectively.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


