Transcript-specific enrichment enables profiling of rare cell states via single-cell RNA sequencing
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
Single-cell genomics technologies have accelerated our understanding of cell-state heterogeneity in diverse contexts. Although single-cell RNA sequencing identifies rare populations that express specific marker transcript combinations, traditional flow sorting requires cell surface markers with high-fidelity antibodies, limiting our ability to interrogate these populations. In addition, many single-cell studies require the isolation of nuclei from tissue, eliminating the ability to enrich learned rare cell states based on extranuclear protein markers. In the present report, we addressed these limitations by developing Programmable Enrichment via RNA FlowFISH by sequencing (PERFF-seq), a scalable assay that enables scRNA-seq profiling of subpopulations defined by the abundance of specific RNA transcripts. Across immune populations (n = 184,126 cells) and fresh-frozen and formalin-fixed, paraffin-embedded brain tissue (n = 33,145 nuclei), we demonstrated that programmable sorting logic via RNA-based cytometry can isolate rare cell populations and uncover phenotypic heterogeneity via downstream, high-throughput, single-cell genomics analyses. Programmable Enrichment via RNA FlowFISH by sequencing (PERFF-seq) isolates rare cells based on RNA marker transcripts for single-cell RNA sequencing profiling of complex tissues, with applicability to a broad variety of samples and cell types.

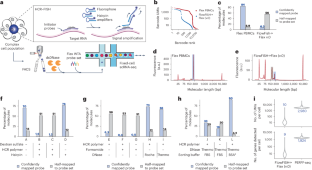
转录特异性富集可以通过单细胞RNA测序来分析罕见的细胞状态
单细胞基因组学技术加速了我们对不同环境下细胞状态异质性的理解。虽然单细胞RNA测序鉴定了表达特定标记转录组合的罕见群体,但传统的流式分选需要具有高保真抗体的细胞表面标记,这限制了我们询问这些群体的能力。此外,许多单细胞研究需要从组织中分离细胞核,从而消除了基于核外蛋白标记丰富已知罕见细胞状态的能力。在本报告中,我们通过通过测序(PERFF-seq)开发可编程富集RNA FlowFISH来解决这些限制,PERFF-seq是一种可扩展的分析方法,可以通过特定RNA转录物的丰度来定义scRNA-seq分析亚群。在免疫群体(n = 184,126个细胞)和新鲜冷冻和福尔马林固定的石蜡包埋脑组织(n = 33,145个细胞核)中,我们证明了基于rna的细胞术的可编程分类逻辑可以分离稀有细胞群体,并通过下游高通量单细胞基因组学分析揭示表型异质性。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


