Determining structures of RNA conformers using AFM and deep neural networks
IF 48.5
1区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 0
Abstract
Much of the human genome is transcribed into RNAs1, many of which contain structural elements that are important for their function. Such RNA molecules—including those that are structured and well-folded2—are conformationally heterogeneous and flexible, which is a prerequisite for function3,4, but this limits the applicability of methods such as NMR, crystallography and cryo-electron microscopy for structure elucidation. Moreover, owing to the lack of a large RNA structure database, and no clear correlation between sequence and structure, approaches such as AlphaFold5 for protein structure prediction do not apply to RNA. Therefore, determining the structures of heterogeneous RNAs remains an unmet challenge. Here we report holistic RNA structure determination method using atomic force microscopy, unsupervised machine learning and deep neural networks (HORNET), a novel method for determining three-dimensional topological structures of RNA using atomic force microscopy images of individual molecules in solution. Owing to the high signal-to-noise ratio of atomic force microscopy, this method is ideal for capturing structures of large RNA molecules in distinct conformations. In addition to six benchmark cases, we demonstrate the utility of HORNET by determining multiple heterogeneous structures of RNase P RNA and the HIV-1 Rev response element (RRE) RNA. Thus, our method addresses one of the major challenges in determining heterogeneous structures of large and flexible RNA molecules, and contributes to the fundamental understanding of RNA structural biology. HORNET, a method that uses unsupervised machine learning and deep neural networks to analyse atomic force microscopy data enables structural determination of RNA molecules in multiple conformations.

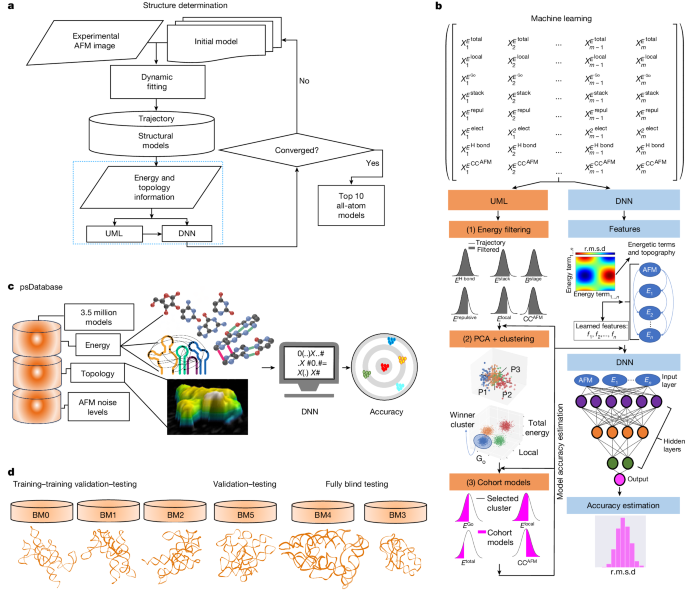
利用原子力显微镜和深度神经网络确定 RNA 构象的结构
人类基因组的大部分被转录成RNAs1,其中许多包含对其功能很重要的结构元素。这类RNA分子——包括那些结构良好且折叠良好的RNA分子——构象不均匀且灵活,这是功能的先决条件3,4,但这限制了核磁共振、晶体学和低温电子显微镜等方法在结构解析方面的适用性。此外,由于缺乏大型RNA结构数据库,序列与结构之间没有明确的相关性,因此诸如AlphaFold5等蛋白质结构预测方法并不适用于RNA。因此,确定异质rna的结构仍然是一个未解决的挑战。在这里,我们报告了利用原子力显微镜、无监督机器学习和深度神经网络(HORNET)的整体RNA结构测定方法,这是一种利用溶液中单个分子的原子力显微镜图像确定RNA三维拓扑结构的新方法。由于原子力显微镜的高信噪比,这种方法是捕获不同构象的大RNA分子结构的理想方法。除了六个基准案例外,我们还通过确定RNase P RNA和HIV-1 Rev应答元件(RRE) RNA的多种异质结构来证明HORNET的实用性。因此,我们的方法解决了确定大而灵活的RNA分子异质结构的主要挑战之一,并有助于对RNA结构生物学的基本理解。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature
综合性期刊-综合性期刊
CiteScore
90.00
自引率
1.20%
发文量
3652
审稿时长
3 months
期刊介绍:
Nature is a prestigious international journal that publishes peer-reviewed research in various scientific and technological fields. The selection of articles is based on criteria such as originality, importance, interdisciplinary relevance, timeliness, accessibility, elegance, and surprising conclusions. In addition to showcasing significant scientific advances, Nature delivers rapid, authoritative, insightful news, and interpretation of current and upcoming trends impacting science, scientists, and the broader public. The journal serves a dual purpose: firstly, to promptly share noteworthy scientific advances and foster discussions among scientists, and secondly, to ensure the swift dissemination of scientific results globally, emphasizing their significance for knowledge, culture, and daily life.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


