Pan-cancer analysis of biallelic inactivation in tumor suppressor genes identifies KEAP1 zygosity as a predictive biomarker in lung cancer
IF 45.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
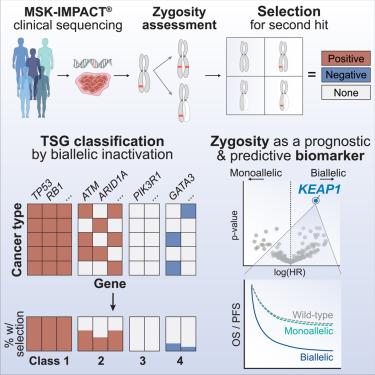
The canonical model of tumor suppressor gene (TSG)-mediated oncogenesis posits that loss of both alleles is necessary for inactivation. Here, through allele-specific analysis of sequencing data from 48,179 cancer patients, we define the prevalence, selective pressure for, and functional consequences of biallelic inactivation across TSGs. TSGs largely assort into distinct classes associated with either pan-cancer (Class 1) or lineage-specific (Class 2) patterns of selection for biallelic loss, although some TSGs are predominantly monoallelically inactivated (Class 3/4). We demonstrate that selection for biallelic inactivation can be utilized to identify driver genes in non-canonical contexts, including among variants of unknown significance (VUSs) of several TSGs such as KEAP1. Genomic, functional, and clinical data collectively indicate that KEAP1 VUSs phenocopy established KEAP1 oncogenic alleles and that zygosity, rather than variant classification, is predictive of therapeutic response. TSG zygosity is therefore a fundamental determinant of disease etiology and therapeutic sensitivity.
求助全文
约1分钟内获得全文
求助全文
来源期刊
Cell
生物-生化与分子生物学
CiteScore
110.00
自引率
0.80%
发文量
396
审稿时长
2 months
期刊介绍:
Cells is an international, peer-reviewed, open access journal that focuses on cell biology, molecular biology, and biophysics. It is affiliated with several societies, including the Spanish Society for Biochemistry and Molecular Biology (SEBBM), Nordic Autophagy Society (NAS), Spanish Society of Hematology and Hemotherapy (SEHH), and Society for Regenerative Medicine (Russian Federation) (RPO).
The journal publishes research findings of significant importance in various areas of experimental biology, such as cell biology, molecular biology, neuroscience, immunology, virology, microbiology, cancer, human genetics, systems biology, signaling, and disease mechanisms and therapeutics. The primary criterion for considering papers is whether the results contribute to significant conceptual advances or raise thought-provoking questions and hypotheses related to interesting and important biological inquiries.
In addition to primary research articles presented in four formats, Cells also features review and opinion articles in its "leading edge" section, discussing recent research advancements and topics of interest to its wide readership.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


