Transposable element exonization generates a reservoir of evolving and functional protein isoforms
IF 45.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
Alternative splicing enhances protein diversity in different ways, including through exonization of transposable elements (TEs). Recent transcriptomic analyses identified thousands of unannotated spliced transcripts with exonizing TEs, but their contribution to the proteome and biological relevance remains unclear. Here, we use transcriptome assembly, ribosome profiling, and proteomics to describe a population of 1,227 unannotated TE exonizing isoforms generated by mRNA splicing and recurrent in human populations. Despite being shorter and lowly expressed, these isoforms are shared between individuals and efficiently translated. Functional analyses show stable expression, specific cellular localization, and, in some cases, modified functions. Exonized TEs are rich in ancient genes, whereas the involved splice sites are recent and can be evolutionarily conserved. In addition, exonized TEs contribute to the secondary structure of the emerging isoforms, supporting their functional relevance. We conclude that TE-spliced isoforms represent a diversity reservoir of functional proteins on which natural selection can act.
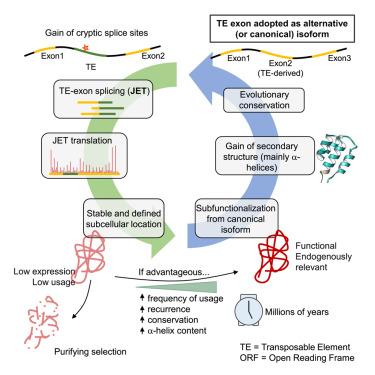
转座因子外显子产生了一个进化的和功能性的蛋白质异构体库
选择性剪接以不同的方式增强蛋白质多样性,包括通过转座因子(te)的外显子化。最近的转录组学分析发现了数千个具有外显子te的未注释剪接转录物,但它们对蛋白质组学和生物学相关性的贡献尚不清楚。在这里,我们使用转录组组装、核糖体分析和蛋白质组学来描述由mRNA剪接产生的1,227个无注释的TE外显异构体,这些异构体在人群中反复出现。尽管这些同工异构体较短且表达较低,但它们在个体之间是共享的,并且可以有效地翻译。功能分析显示稳定的表达,特定的细胞定位,在某些情况下,修改功能。外显子te富含古代基因,而所涉及的剪接位点是最近的,可能是进化保守的。此外,外显子化的te有助于新出现的同工异构体的二级结构,支持其功能相关性。我们得出结论,te剪接异构体代表了自然选择可以发挥作用的功能蛋白的多样性储存库。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Cell
生物-生化与分子生物学
CiteScore
110.00
自引率
0.80%
发文量
396
审稿时长
2 months
期刊介绍:
Cells is an international, peer-reviewed, open access journal that focuses on cell biology, molecular biology, and biophysics. It is affiliated with several societies, including the Spanish Society for Biochemistry and Molecular Biology (SEBBM), Nordic Autophagy Society (NAS), Spanish Society of Hematology and Hemotherapy (SEHH), and Society for Regenerative Medicine (Russian Federation) (RPO).
The journal publishes research findings of significant importance in various areas of experimental biology, such as cell biology, molecular biology, neuroscience, immunology, virology, microbiology, cancer, human genetics, systems biology, signaling, and disease mechanisms and therapeutics. The primary criterion for considering papers is whether the results contribute to significant conceptual advances or raise thought-provoking questions and hypotheses related to interesting and important biological inquiries.
In addition to primary research articles presented in four formats, Cells also features review and opinion articles in its "leading edge" section, discussing recent research advancements and topics of interest to its wide readership.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


