Trace_y: Software algorithms for structural analysis of individual helical filaments by three-dimensional contact point reconstruction atomic force microscopy
IF 4.4
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
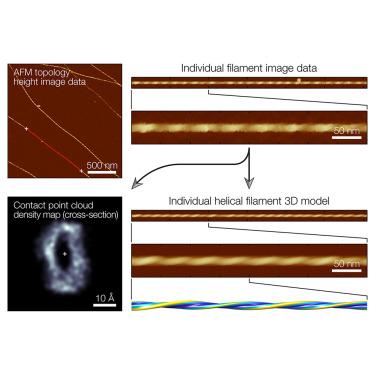
Atomic force microscopy (AFM) is a powerful and increasingly accessible technology that has a wide range of bio-imaging applications. AFM is capable of producing detailed three-dimensional topographical images with high signal-to-noise ratio, which enables the structural features of individual molecules to be studied without the need for ensemble averaging. Here, a software tool Trace_y, designed to reconstruct the three-dimensional surface envelopes of individual helical filament structures from topographical AFM images, is presented. Workflow using Trace_y is demonstrated on the structural analysis of individual helical amyloid protein fibrils where the assembly mechanism of heterogeneous, complex and diverse fibril populations due to structural polymorphism is not understood. The algorithms presented here allow structural information encoded in topographical AFM height images to be extracted and understood as three-dimensional (3D) contact point clouds. This approach will facilitate the use of AFM in structural biology to understand molecular structures and behaviors at individual molecule level.
用三维接触点重建原子力显微镜对单个螺旋细丝进行结构分析的软件算法
原子力显微镜(AFM)是一种功能强大且日益普及的技术,具有广泛的生物成像应用。AFM能够产生具有高信噪比的详细的三维地形图像,这使得无需集合平均即可研究单个分子的结构特征。本文介绍了一种软件工具Trace_y,用于从地形AFM图像中重建单个螺旋细丝结构的三维表面包络。使用Trace_y对单个螺旋淀粉样蛋白原纤维的结构分析进行了工作流演示,其中由于结构多态性而导致的异质,复杂和多样化的原纤维群体的组装机制尚不清楚。本文提出的算法允许在地形AFM高度图像中编码的结构信息被提取并理解为三维(3D)接触点云。这种方法将促进AFM在结构生物学中的应用,以在单个分子水平上理解分子结构和行为。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


