Coastal influence on microbiomes of the Southwest Atlantic Ocean
引用次数: 0
Abstract
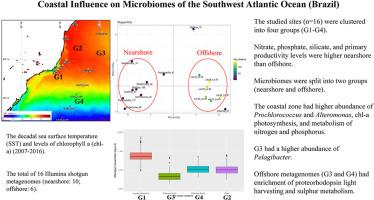
The possible influence of the continent on the microbiome of the Southwest Atlantic Ocean (SAO) is not yet fully known. The aim of the present study was to evaluate the metagenomic diversity of the SAO nearshore in different locations (e.g., rivers and upwellings) and offshore locations. The studied sites (n = 16) were clustered into four groups (G1-G4) according to the decadal sea surface temperature (SST) and levels of chlorophyll a (chl-a) (2007–2016). G1 (coastal upwelling) presented the highest chl-a values, G2 (shelf) had the highest SST, G3 (offshore north) had the lowest chl-a levels, and G4 (offshore south) was the coldest site. The higher nutrient levels may contribute to higher chl-a-based primary productivity and possibly higher bacterial productivity nearshore. The taxonomic and functional diversity of the total of 16 Illumina shotgun metagenomes were analysed (nearshore: 10, offshore: 6; mean ± standard deviation: 1.87 ± 0.41 million reads per sample). The SAO microbiomes were split into two groups: nearshore and offshore. The SAO coastal zone had higher abundance of picoplanktonic cyanobacteria (e.g., Prochlorococcus), copiotrophic bacteria (e.g., Alteromonas), chl-a photosynthesis, and metabolism of nitrogen and phosphorus. In clear contrast with offshore microbiomes, nearshore microbiomes were heterogenous as a possible effect of riverine runoff and the upwelling. G1 microbiomes had higher abundance of copiotrophic bacteria, while G2 had a higher abundance of Prochlorococcus, and G3 had a higher abundance of Pelagibacter. Offshore metagenomes (G3 and G4) had enrichment of proteorhodopsin light harvesting and sulfur metabolism, possible relevant processes in the SAO. Metagenome assembled genomes (MAGs) demonstrated significant novel microbiome diversity. The most abundant MAGs belong to Flavobacteria and Euryarchaeota. The discrimination of microbial populations and metabolisms in the four studied oceanographic groups provides a first glimpse on the microbial landscape in the SAO.
西南大西洋沿岸对微生物组的影响
非洲大陆对西南大西洋(SAO)微生物群可能产生的影响尚不完全清楚。本研究的目的是评估SAO近岸不同位置(如河流和上升流)和近海位置的宏基因组多样性。根据2007-2016年的年代际海表温度(SST)和叶绿素a (chl-a)水平,将16个研究点分为G1-G4组。G1(沿海上升流)的chl-a最高,G2(陆架)的海温最高,G3(近海北部)的chl-a最低,G4(近海南部)的chl-a最冷。较高的营养水平可能有助于提高以chl为基础的初级生产力,并可能提高近岸的细菌生产力。对16个Illumina shotgun元基因组进行了分类和功能多样性分析(近岸:10个,近海:6个;平均值±标准差:1.87±0.41万reads / sample)。SAO微生物组被分为两组:近岸和近海。SAO海岸带具有较高的微浮游蓝藻(如原绿球藻)、共生细菌(如互生单胞菌)、chl-a光合作用和氮磷代谢丰度。与近海微生物群落形成鲜明对比的是,近岸微生物群落具有异质性,这可能是河流径流和上升流的影响。G1菌群中coiotrophic bacteria丰度较高,G2菌群中原绿球藻丰度较高,G3菌群中Pelagibacter丰度较高。离岸宏基因组(G3和G4)富含蛋白紫质光收集和硫代谢,可能与SAO有关。宏基因组组装基因组(MAGs)显示出显著的新微生物组多样性。最丰富的MAGs属于黄杆菌门和Euryarchaeota。在四个研究的海洋学组中对微生物种群和代谢的区分提供了对SAO微生物景观的初步了解。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文

 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


