Single-cell multiomics analysis reveals dynamic clonal evolution and targetable phenotypes in acute myeloid leukemia with complex karyotype
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
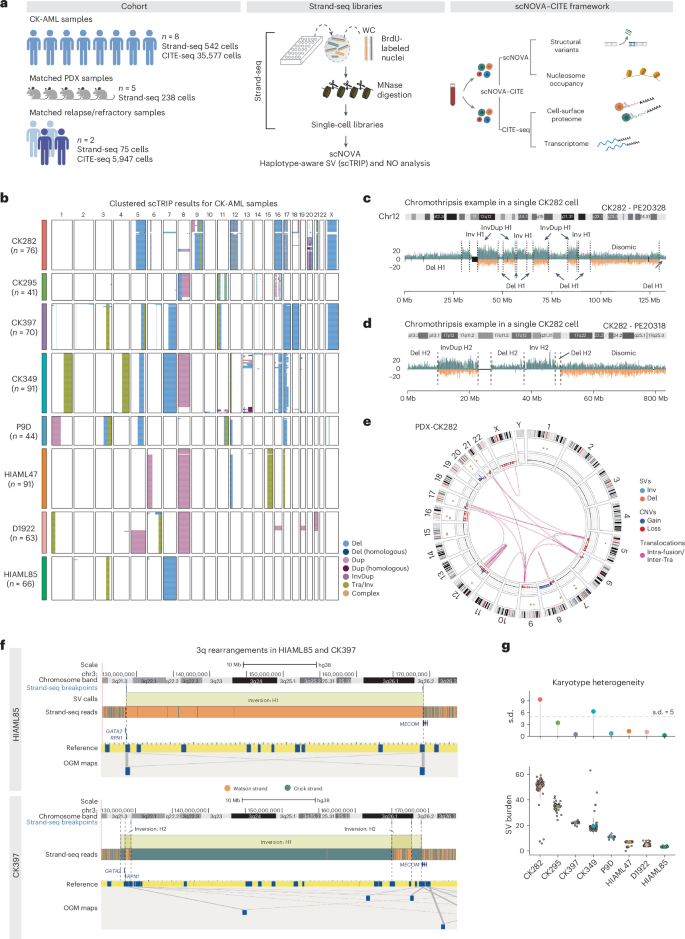
Chromosomal instability is a major driver of intratumoral heterogeneity (ITH), promoting tumor progression. In the present study, we combined structural variant discovery and nucleosome occupancy profiling with transcriptomic and immunophenotypic changes in single cells to study ITH in complex karyotype acute myeloid leukemia (CK-AML). We observed complex structural variant landscapes within individual cells of patients with CK-AML characterized by linear and circular breakage–fusion–bridge cycles and chromothripsis. We identified three clonal evolution patterns in diagnosis or salvage CK-AML (monoclonal, linear and branched polyclonal), with 75% harboring multiple subclones that frequently displayed ongoing karyotype remodeling. Using patient-derived xenografts, we demonstrated varied clonal evolution of leukemic stem cells (LSCs) and further dissected subclone-specific drug–response profiles to identify LSC-targeting therapies, including BCL-xL inhibition. In paired longitudinal patient samples, we further revealed genetic evolution and cell-type plasticity as mechanisms of disease progression. By dissecting dynamic genomic, phenotypic and functional complexity of CK-AML, our findings offer clinically relevant avenues for characterizing and targeting disease-driving LSCs. An integrated single-cell multiomic analysis of complex karyotype acute myeloid leukemia characterizes intratumoral heterogeneity and highlights links to therapeutic sensitivities.

单细胞多组学分析揭示复杂核型急性髓性白血病的动态克隆进化和靶向表型
染色体不稳定性是瘤内异质性(ITH)的主要驱动因素,可促进肿瘤进展。在本研究中,我们将结构变异的发现和核小体占位谱分析与单细胞的转录组学和免疫表型变化相结合,研究了复杂核型急性髓性白血病(CK-AML)中的ITH。我们在 CK-AML 患者的单个细胞中观察到了复杂的结构变异景观,其特征是线性和环状断裂-融合-桥接循环以及染色体三分裂。我们在确诊或抢救的 CK-AML 患者中发现了三种克隆进化模式(单克隆、线性和分支多克隆),其中 75% 的患者携带多个亚克隆,这些亚克隆经常表现出持续的核型重塑。我们利用源自患者的异种移植,证实了白血病干细胞(LSC)的不同克隆演变,并进一步剖析了亚克隆特异性药物反应谱,以确定针对LSC的疗法,包括BCL-xL抑制剂。在成对的纵向患者样本中,我们进一步揭示了基因进化和细胞类型可塑性作为疾病进展机制的作用。通过剖析CK-AML的动态基因组、表型和功能复杂性,我们的研究结果为描述和靶向疾病驱动的LSC提供了临床相关的途径。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


