Microbial communities in slow sand filters for drinking water treatment adapt to organic matter altered by ozonation
IF 12.4
1区 环境科学与生态学
Q1 ENGINEERING, ENVIRONMENTAL
引用次数: 0
Abstract
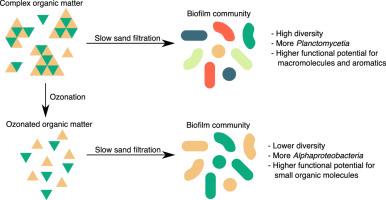
Changing natural organic matter quality from anthropogenic activity and stricter requirements for micropollutant removal challenges existing systems for drinking water production. Ozonation of water followed by biofiltration, such as passage through a slow sand filter (SSF), is a partial solution. Biofiltration relies on biofilms (microbial communities within extracellular matrices). However, the effects of ozonation on SSF microbial communities are unknown. In this study, genome-resolved and read-based metagenomics were used to compare the microbial communities of two full-scale SSFs employing conventional pre-treatment to a 20 m2 SSF operated in parallel with ozonation as additional pre-treatment.
The SSF microbial community receiving ozonated water was less diverse than those receiving non-ozonated water. Families Hyphomicrobiaceae, Acetobacteraceae, Sphingomonadaceae and Burkholderiaceae were more abundant when ozone was used, as were genes for metabolism of single-carbon organic compounds. Conversely, genes for metabolism of aromatic compounds and fatty acids were less abundant. Metagenome assembled genomes associated with the non-ozonated SSFs were enriched with several glycoside hydrolases, while those associated with the ozonated SSF were enriched with genes for 1-2 carbon compound metabolism. No indications of increased microbial risk (pathogens or antibiotic resistance genes) were detected as a consequence of ozonation.
This study shows how microbial communities of SSFs adapt to changes in organic matter quality, highlighting the key role of biofilters for production of safe and sustainable drinking water in a changing climate.
用于饮用水处理的慢沙过滤器中的微生物群落适应臭氧改变的有机物
人类活动导致天然有机物质量不断变化,对去除微污染物的要求也更加严格,这对现有的饮用水生产系统提出了挑战。对水进行臭氧处理,然后进行生物过滤,如通过慢沙过滤器(SSF),是一种部分解决方案。生物过滤依赖于生物膜(细胞外基质中的微生物群落)。然而,臭氧对 SSF 微生物群落的影响尚不清楚。本研究采用基因组分辨和基于读数的元基因组学方法,比较了两个采用传统预处理方法的全规模固态沉淀池的微生物群落,以及一个与臭氧作为附加预处理方法并行运行的 20 平方米固态沉淀池的微生物群落。使用臭氧时,Hyphomicrobiaceae、Acetobacteraceae、Sphingomonadaceae 和 Burkholderiaceae 家族的数量更多,代谢单碳有机化合物的基因也更多。相反,代谢芳香族化合物和脂肪酸的基因则较少。与非臭氧 SSF 相关的元基因组中富含多种糖苷水解酶,而与臭氧 SSF 相关的元基因组中富含 1-2 碳化合物代谢基因。这项研究显示了 SSF 微生物群落如何适应有机物质量的变化,突出了生物过滤器在不断变化的气候条件下生产安全、可持续饮用水方面的关键作用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Water Research
环境科学-工程:环境
CiteScore
20.80
自引率
9.40%
发文量
1307
审稿时长
38 days
期刊介绍:
Water Research, along with its open access companion journal Water Research X, serves as a platform for publishing original research papers covering various aspects of the science and technology related to the anthropogenic water cycle, water quality, and its management worldwide. The audience targeted by the journal comprises biologists, chemical engineers, chemists, civil engineers, environmental engineers, limnologists, and microbiologists. The scope of the journal include:
•Treatment processes for water and wastewaters (municipal, agricultural, industrial, and on-site treatment), including resource recovery and residuals management;
•Urban hydrology including sewer systems, stormwater management, and green infrastructure;
•Drinking water treatment and distribution;
•Potable and non-potable water reuse;
•Sanitation, public health, and risk assessment;
•Anaerobic digestion, solid and hazardous waste management, including source characterization and the effects and control of leachates and gaseous emissions;
•Contaminants (chemical, microbial, anthropogenic particles such as nanoparticles or microplastics) and related water quality sensing, monitoring, fate, and assessment;
•Anthropogenic impacts on inland, tidal, coastal and urban waters, focusing on surface and ground waters, and point and non-point sources of pollution;
•Environmental restoration, linked to surface water, groundwater and groundwater remediation;
•Analysis of the interfaces between sediments and water, and between water and atmosphere, focusing specifically on anthropogenic impacts;
•Mathematical modelling, systems analysis, machine learning, and beneficial use of big data related to the anthropogenic water cycle;
•Socio-economic, policy, and regulations studies.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


