The kinetoplastid kinetochore protein KKT23 acetyltransferase is a structural homolog of GCN5 that acetylates the histone H2A C-terminal tail
IF 4.4
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
The kinetochore is the macromolecular protein machine that drives chromosome segregation in eukaryotes. In an evolutionarily divergent group of organisms called kinetoplastids, kinetochores are built using a unique set of proteins (KKT1–25 and KKIP1–12). KKT23 is a constitutively localized kinetochore protein containing a C-terminal acetyltransferase domain of unknown function. Here, using X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy, we have determined the structure and dynamics of the KKT23 acetyltransferase domain from Trypanosoma brucei and found that it is structurally similar to the GCN5 histone acetyltransferase domain. We find that KKT23 can acetylate the C-terminal tail of histone H2A and that knockdown of KKT23 results in decreased H2A acetylation levels in T. brucei. Finally, we have determined the crystal structure of the N-terminal region of KKT23 and shown that it interacts with KKT22. Our study provides important insights into the structure and function of the unique kinetochore acetyltransferase in trypanosomes.
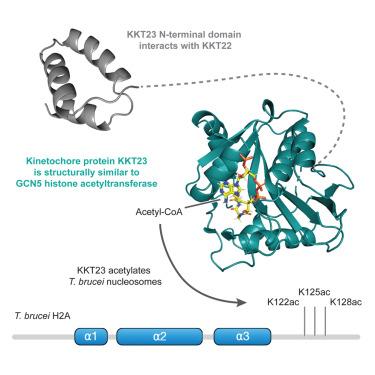
核原生动物动点蛋白 KKT23 乙酰基转移酶是 GCN5 的结构同源物,它能使组蛋白 H2A C 端尾乙酰化
动核是真核生物中驱动染色体分离的大分子蛋白质机器。在进化过程中出现分化的一类生物(称为动点细胞)中,动点由一组独特的蛋白质(KKT1-25 和 KKIP1-12)构建。KKT23 是一种组成型定位的动点核蛋白,含有一个功能未知的 C 端乙酰转移酶结构域。在这里,我们利用 X 射线晶体学和核磁共振(NMR)光谱测定了布氏锥虫 KKT23 乙酰转移酶结构域的结构和动力学,发现它在结构上与 GCN5 组蛋白乙酰转移酶结构域相似。我们发现 KKT23 能使组蛋白 H2A 的 C 端尾乙酰化,而且敲除 KKT23 会导致布氏锥虫 H2A 乙酰化水平下降。最后,我们测定了 KKT23 N 端区域的晶体结构,并证明它与 KKT22 相互作用。我们的研究对锥虫中独特的动点核乙酰转移酶的结构和功能提供了重要的见解。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


