Omics-centric evidences of fipronil biodegradation by Rhodococcus sp. FIP_B3.
IF 7.6
2区 环境科学与生态学
Q1 ENVIRONMENTAL SCIENCES
引用次数: 0
Abstract
The widespread use of the pesticide fipronil in domestic and agriculture sectors has resulted in its accumulation across the environment. Its use to assure food security has inadvertently affected soil microbiome composition, fertility and, ultimately, human health. Degradation of residual fipronil present in the environment using specific microbial species is a promising strategy for its removal. The present study delves into the omics approach for fipronil biodegradation using the native bacterium Rhodococcus sp. FIP B3. It has been observed that within 40 days, nearly 84% of the insecticide gets degraded. The biodegradation follows a pseudo-first-order kinetics (k = 0.0197/d with a half-life of ∼11 days). Whole genome analysis revealed Cytochrome P450 monooxygenase, peroxidase-related enzyme, haloalkane dehalogenase, 2-nitropropane dioxygenase, and aconitate hydratase are involved in the degradation process. Fipronil-sulfone, 5-amino-1-(2-chloro-4-(trifluoromethyl)phenyl)-4- ((trifluoromethyl)sulfonyl)-1H-pyrazole-3-carbonitrile, (E)-5-chloro-2-oxo-3- (trifluoromethyl)pent-4-enoic acid, 4,4,4-trifluoro-2-oxobutanoic acid, and 3,3,3- trifluoropropanoic acid were identified as the major metabolites that support the bacterial degradation of fipronil. In-silico molecular docking and molecular dynamic simulation-based analyses of degradation pathway intermediates with their respective enzymes have indicated stable interactions with significant binding energies (-5.9 to -9.7 kcal/mol). These results have provided the mechanistic cause of the elevated potential of Rhodococcus sp. FIP_B3 for fipronil degradation and will be advantageous in framing appropriate strategies for the bioremediation of fipronil-contaminated environment.
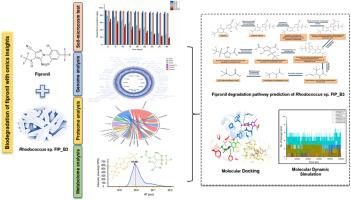
Rhodococcus sp. FIP_B3 对氟虫腈生物降解的 Omics 中心证据。
杀虫剂氟虫腈在家庭和农业部门的广泛使用已导致其在整个环境中的积累。为确保粮食安全而使用氟虫腈无意中影响了土壤微生物组的组成、肥力,并最终影响了人类健康。利用特定微生物物种降解环境中的残留氟虫腈是一种很有前景的除氟策略。本研究利用本地细菌 Rhodococcus sp. FIP B3 对氟虫腈的生物降解进行了全方位研究。据观察,在 40 天内,近 84% 的杀虫剂被降解。生物降解遵循假一阶动力学(k = 0.0197/d,半衰期为 11 天)。全基因组分析表明,细胞色素 P450 单加氧酶、过氧化物酶相关酶、卤代烷烃脱卤酶、2-硝基丙烷二加氧酶和乌头加氢酶参与了降解过程。氟虫腈砜,5-氨基-1-(2-氯-4-(三氟甲基)苯基)-4-((三氟甲基)磺酰基)-1H-吡唑-3-甲腈,(E)-5-氯-2-氧代-3-(三氟甲基)戊-4-烯酸、经鉴定,4,4,4-三氟-2-氧代丁酸和 3,3,3- 三氟丙酸是支持细菌降解氟虫腈的主要代谢产物。对降解途径中间产物与各自酶进行的室内分子对接和分子动态模拟分析表明,它们之间存在稳定的相互作用,且结合能显著(-5.9 至 -9.7 kcal/mol)。这些结果从机理上说明了 Rhodococcus sp. FIP_B3 在降解氟虫腈方面潜力巨大的原因,并将有助于为氟虫腈污染环境的生物修复制定适当的策略。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Environmental Pollution
环境科学-环境科学
CiteScore
16.00
自引率
6.70%
发文量
2082
审稿时长
2.9 months
期刊介绍:
Environmental Pollution is an international peer-reviewed journal that publishes high-quality research papers and review articles covering all aspects of environmental pollution and its impacts on ecosystems and human health.
Subject areas include, but are not limited to:
• Sources and occurrences of pollutants that are clearly defined and measured in environmental compartments, food and food-related items, and human bodies;
• Interlinks between contaminant exposure and biological, ecological, and human health effects, including those of climate change;
• Contaminants of emerging concerns (including but not limited to antibiotic resistant microorganisms or genes, microplastics/nanoplastics, electronic wastes, light, and noise) and/or their biological, ecological, or human health effects;
• Laboratory and field studies on the remediation/mitigation of environmental pollution via new techniques and with clear links to biological, ecological, or human health effects;
• Modeling of pollution processes, patterns, or trends that is of clear environmental and/or human health interest;
• New techniques that measure and examine environmental occurrences, transport, behavior, and effects of pollutants within the environment or the laboratory, provided that they can be clearly used to address problems within regional or global environmental compartments.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


