Versatile xylose and arabinose genetic switches development for yeasts
IF 6.8
1区 生物学
Q1 BIOTECHNOLOGY & APPLIED MICROBIOLOGY
引用次数: 0
Abstract
Inducible transcription systems are essential tools in genetic engineering, where tight control, strong inducibility and fast response with cost-effective inducers are highly desired. However, existing systems in yeasts are rarely used in large-scale fermentations due to either cost-prohibitive inducers or incompatible performance. Here, we developed powerful xylose and arabinose induction systems in Saccharomyces cerevisiae, utilizing eukaryotic activators XlnR and AraRA from Aspergillus species and bacterial repressors XylR and AraRR. By integrating these signals into a highly-structured synthetic promoter, we created dual-mode systems with strong outputs and minimal leakiness. These systems demonstrated over 4000- and 300-fold regulation with strong activation and rapid response. The dual-mode xylose system was fully activated by xylose-rich agricultural residues like corncob hydrolysate, outperforming existing systems in terms of leakiness, inducibility, dynamic range, induction rate, and growth impact on host. We validated their utility in metabolic engineering with high-titer linalool production and demonstrated the transferability of the XlnR-based xylose induction system to Pichia pastoris, Candida glabrata and Candida albicans. This work provides robust genetic switches for yeasts and a general strategy for integrating activation-repression signals into synthetic promoters to achieve optimal performance.
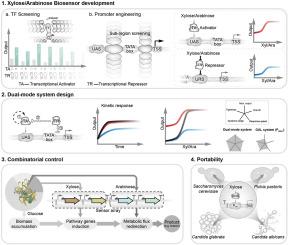
为酵母开发多功能木糖和阿拉伯糖基因开关。
诱导转录系统是基因工程中必不可少的工具,在这种系统中,人们非常需要严格的控制、强大的诱导性和快速的反应,同时还需要具有成本效益的诱导剂。然而,由于诱导剂成本过高或性能不兼容,现有的酵母系统很少用于大规模发酵。在这里,我们利用来自曲霉的真核激活剂 XlnR 和 AraRA 以及细菌抑制剂 XylR 和 AraRR,在酿酒酵母中开发出了强大的木糖和阿拉伯糖诱导系统。通过将这些信号整合到高度结构化的合成启动子中,我们创建了具有强大输出和最小泄漏的双模式系统。这些系统的调控能力分别超过 4000 倍和 300 倍,并且具有很强的激活能力和快速反应能力。富含木糖的农业残留物(如玉米芯水解物)可完全激活双模式木糖系统,该系统在泄漏性、诱导性、动态范围、诱导率以及对宿主生长的影响方面均优于现有系统。我们通过高滴度芳樟醇的生产验证了它们在代谢工程中的实用性,并证明了基于 XlnR 的木糖诱导系统在 Pichia pastoris、Candida glabrata 和 Candida albicans 中的可移植性。这项工作为酵母提供了强大的基因开关,并为将激活-抑制信号整合到合成启动子中以实现最佳性能提供了通用策略。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Metabolic engineering
工程技术-生物工程与应用微生物
CiteScore
15.60
自引率
6.00%
发文量
140
审稿时长
44 days
期刊介绍:
Metabolic Engineering (MBE) is a journal that focuses on publishing original research papers on the directed modulation of metabolic pathways for metabolite overproduction or the enhancement of cellular properties. It welcomes papers that describe the engineering of native pathways and the synthesis of heterologous pathways to convert microorganisms into microbial cell factories. The journal covers experimental, computational, and modeling approaches for understanding metabolic pathways and manipulating them through genetic, media, or environmental means. Effective exploration of metabolic pathways necessitates the use of molecular biology and biochemistry methods, as well as engineering techniques for modeling and data analysis. MBE serves as a platform for interdisciplinary research in fields such as biochemistry, molecular biology, applied microbiology, cellular physiology, cellular nutrition in health and disease, and biochemical engineering. The journal publishes various types of papers, including original research papers and review papers. It is indexed and abstracted in databases such as Scopus, Embase, EMBiology, Current Contents - Life Sciences and Clinical Medicine, Science Citation Index, PubMed/Medline, CAS and Biotechnology Citation Index.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


