Decoding transcriptional identity in developing human sensory neurons and organoid modeling
IF 45.5
1区 生物学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
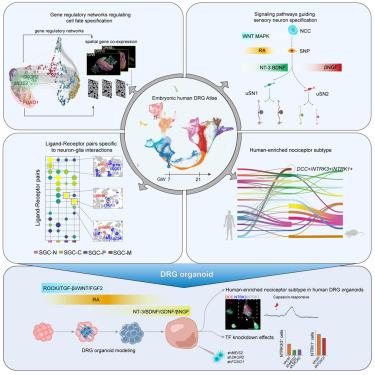
Dorsal root ganglia (DRGs) play a crucial role in processing sensory information, making it essential to understand their development. Here, we construct a single-cell spatiotemporal transcriptomic atlas of human embryonic DRG. This atlas reveals the diversity of cell types and highlights the extrinsic signaling cascades and intrinsic regulatory hierarchies that guide cell fate decisions, including neuronal/glial lineage restriction, sensory neuron differentiation and specification, and the formation of neuron-satellite glial cell (SGC) units. Additionally, we identify a human-enriched NTRK3+/DCC+ nociceptor subtype, which is involved in multimodal nociceptive processing. Mimicking the programmed activation of signaling pathways in vivo, we successfully establish functional human DRG organoids and underscore the critical roles of transcriptional regulators in the fate commitment of unspecialized sensory neurons (uSNs). Overall, our research elucidates the multilevel signaling pathways and transcription factor (TF) regulatory hierarchies that underpin the diversity of somatosensory neurons, emphasizing the phenotypic distinctions in human nociceptor subtypes.
解码发育中人类感觉神经元的转录特征和类器官模型
背根神经节(DRG)在处理感觉信息方面起着至关重要的作用,因此了解其发育过程至关重要。在这里,我们构建了人类胚胎背根神经节的单细胞时空转录组图谱。该图谱揭示了细胞类型的多样性,并突出了指导细胞命运决定的外在信号级联和内在调控层次,包括神经元/神经胶质细胞系的限制、感觉神经元的分化和规格化以及神经元-卫星胶质细胞(SGC)单元的形成。此外,我们还发现了一种人类丰富的 NTRK3+/DCC+ 痛觉感受器亚型,它参与了多模式痛觉处理。模仿体内信号通路的程序化激活,我们成功地建立了功能性人DRG器官组织,并强调了转录调节因子在非特化感觉神经元(uSNs)命运承诺中的关键作用。总之,我们的研究阐明了支撑躯体感觉神经元多样性的多级信号通路和转录因子(TF)调控层次,强调了人类痛觉感受器亚型的表型差异。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Cell
生物-生化与分子生物学
CiteScore
110.00
自引率
0.80%
发文量
396
审稿时长
2 months
期刊介绍:
Cells is an international, peer-reviewed, open access journal that focuses on cell biology, molecular biology, and biophysics. It is affiliated with several societies, including the Spanish Society for Biochemistry and Molecular Biology (SEBBM), Nordic Autophagy Society (NAS), Spanish Society of Hematology and Hemotherapy (SEHH), and Society for Regenerative Medicine (Russian Federation) (RPO).
The journal publishes research findings of significant importance in various areas of experimental biology, such as cell biology, molecular biology, neuroscience, immunology, virology, microbiology, cancer, human genetics, systems biology, signaling, and disease mechanisms and therapeutics. The primary criterion for considering papers is whether the results contribute to significant conceptual advances or raise thought-provoking questions and hypotheses related to interesting and important biological inquiries.
In addition to primary research articles presented in four formats, Cells also features review and opinion articles in its "leading edge" section, discussing recent research advancements and topics of interest to its wide readership.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


