FoxA1/2-dependent epigenomic reprogramming drives lineage switching in lung adenocarcinoma
IF 10.7
1区 生物学
Q1 CELL BIOLOGY
引用次数: 0
Abstract
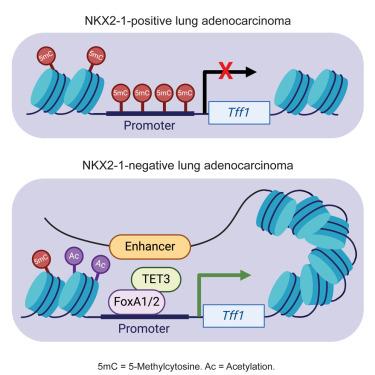
The ability of cancer cells to undergo identity changes (i.e., lineage plasticity) plays a key role in tumor progression and response to therapy. Loss of the pulmonary lineage specifier NKX2-1 in KRAS-driven lung adenocarcinoma (LUAD) enhances tumor progression and causes a FoxA1/2-dependent pulmonary-to-gastric lineage switch. However, the mechanisms by which FoxA1/2 activate a latent gastric identity in the lung remain largely unknown. Here, we show that FoxA1/2 reprogram the epigenetic landscape of gastric-specific genes after NKX2-1 loss in mouse models by facilitating ten-eleven translocation (TET)2/3 recruitment, DNA demethylation, histone 3 lysine 27 acetylation (H3K27ac) deposition, and three-dimensional (3D) chromatin interactions. FoxA1/2-mediated DNA methylation changes are highly conserved in human endodermal development and in progression of human lung and pancreatic neoplasia. Furthermore, oncogenic signaling is required for specific elements of FoxA1/2-dependent epigenetic reprogramming. This work demonstrates the role of FoxA1/2 in rewiring the DNA methylation and 3D chromatin landscape of NKX2-1-negative LUAD to drive cancer cell lineage switching.
FoxA1/2 依赖性表观基因组重编程驱动肺腺癌的系谱转换
癌细胞发生身份变化的能力(即谱系可塑性)在肿瘤进展和治疗反应中起着关键作用。在 KRAS 驱动的肺腺癌(LUAD)中,肺系标志物 NKX2-1 的缺失会增强肿瘤的进展,并导致 FoxA1/2 依赖性的肺系到胃系的转换。然而,FoxA1/2激活肺部潜伏胃特性的机制在很大程度上仍然未知。在这里,我们发现 FoxA1/2 通过促进十-十一易位(TET)2/3 招募、DNA 去甲基化、组蛋白 3 赖氨酸 27 乙酰化(H3K27ac)沉积和三维(3D)染色质相互作用,在小鼠模型中对 NKX2-1 缺失后的胃特异性基因的表观遗传景观进行了重编程。FoxA1/2 介导的 DNA 甲基化变化在人类内胚层发育以及人类肺部和胰腺肿瘤进展过程中高度保守。此外,FoxA1/2依赖的表观遗传重编程的特定元素需要致癌信号。这项工作证明了FoxA1/2在重构NKX2-1阴性LUAD的DNA甲基化和三维染色质图谱以驱动癌细胞谱系转换中的作用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Developmental cell
生物-发育生物学
CiteScore
18.90
自引率
1.70%
发文量
203
审稿时长
3-6 weeks
期刊介绍:
Developmental Cell, established in 2001, is a comprehensive journal that explores a wide range of topics in cell and developmental biology. Our publication encompasses work across various disciplines within biology, with a particular emphasis on investigating the intersections between cell biology, developmental biology, and other related fields. Our primary objective is to present research conducted through a cell biological perspective, addressing the essential mechanisms governing cell function, cellular interactions, and responses to the environment. Moreover, we focus on understanding the collective behavior of cells, culminating in the formation of tissues, organs, and whole organisms, while also investigating the consequences of any malfunctions in these intricate processes.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


