The ZmCPK39–ZmDi19–ZmPR10 immune module regulates quantitative resistance to multiple foliar diseases in maize
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
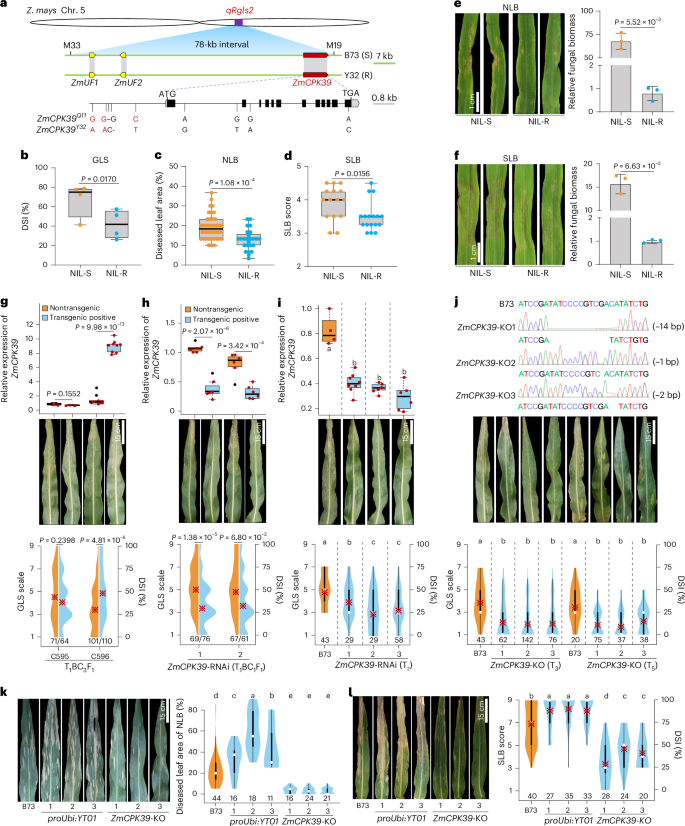
Gray leaf spot, northern leaf blight and southern leaf blight are three of the most destructive foliar diseases affecting maize (Zea mays L.). Here we identified a gene, ZmCPK39, that encodes a calcium-dependent protein kinase and negatively regulates quantitative resistance to these three diseases. The ZmCPK39 allele in the resistant line displayed significantly lower pathogen-induced gene expression than that in the susceptible line. A marked decrease in ZmCPK39 abundance mitigated the phosphorylation and degradation of the transcription factor ZmDi19. This led to elevated expression of ZmPR10, a gene known to encode an antimicrobial protein, thereby enhancing maize resistance to foliar diseases. Moreover, the F1 hybrid with reduced ZmCPK39 expression favored disease resistance, thereby increasing yield. Hence, the discovery of the ZmCPK39–ZmDi19–ZmPR10 immune module provides insight into the mechanisms underlying broad-spectrum quantitative disease resistance and also offers a new avenue for the genetic control of maize foliar diseases. A calcium-dependent protein kinase ZmCPK39 regulates quantitative resistance to multiple foliar diseases in maize through the ZmCPK39–ZmDi19–ZmPR10 immune module.

ZmCPK39-ZmDi19-ZmPR10 免疫模块调控玉米对多种叶面病害的定量抗性
灰叶斑病、北方叶枯病和南方叶枯病是影响玉米(Zea mays L.)的三种最具破坏性的叶部病害。 在这里,我们发现了一个编码钙依赖性蛋白激酶的基因 ZmCPK39,该基因负调控对这三种病害的定量抗性。抗病品系中的 ZmCPK39 等位基因的病原诱导基因表达量明显低于易感品系。ZmCPK39 丰度的显著降低减轻了转录因子 ZmDi19 的磷酸化和降解。这导致 ZmPR10(一种已知编码抗菌蛋白的基因)的表达量增加,从而增强了玉米对叶面病害的抗性。此外,ZmCPK39 表达量减少的 F1 代杂交种抗病性更强,从而提高了产量。因此,ZmCPK39-ZmDi19-ZmPR10 免疫模块的发现有助于深入了解广谱定量抗病性的内在机制,也为玉米叶部病害的遗传控制提供了一条新途径。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


