LEDGF interacts with the NID domain of MeCP2 and modulates MeCP2 condensates
IF 4.4
2区 生物学
Q2 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
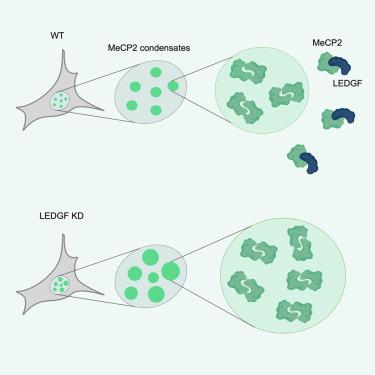
Methyl-CpG-binding protein 2 (MeCP2) is a ubiquitously expressed nuclear protein involved in transcriptional regulation and chromatin remodeling. MeCP2 exists in two isoforms, MeCP2 E1 and E2, which share the same functional domains. Loss-of-function mutations in the MeCP2 gene are the main cause of Rett syndrome (RTT). Previous studies identified a complex formation between MeCP2 and lens epithelium derived growth factor (LEDGF), a transcriptional regulator that exists in two isoforms, LEDGF/p75 and LEDGF/p52. Here, we characterized the molecular and functional interaction between MeCP2 and LEDGF. The NCoR interaction domain (NID) domain in MeCP2 is essential for the direct binding to the PWWP-CR1 region of LEDGF. Introduction of R306C, an RTT mutation in the NID of MeCP2, reduced the interaction with LEDGF. Our data reveal mutual inhibition of MeCP2 and LEDGF multimerization due to overlapping binding sites. Aligning with this observation, LEDGF depletion resulted in larger MeCP2 and DNA foci in NIH3T3 cells, suggesting a role for the MeCP2-LEDGF complex in chromatin organization.
LEDGF与MeCP2的NID结构域相互作用并调节MeCP2的凝结
甲基 CpG 结合蛋白 2(MeCP2)是一种广泛表达的核蛋白,参与转录调控和染色质重塑。MeCP2 有两种异构体,即 MeCP2 E1 和 E2,它们具有相同的功能域。MeCP2 基因的功能缺失突变是导致雷特综合征(RTT)的主要原因。先前的研究发现,MeCP2 与晶状体上皮衍生生长因子(LEDGF)之间形成了复合物,LEDGF 是一种转录调节因子,有两种异构体:LEDGF/p75 和 LEDGF/p52。在这里,我们研究了 MeCP2 与 LEDGF 之间的分子和功能相互作用。MeCP2中的NCoR相互作用结构域(NID)是与LEDGF的PWWP-CR1区域直接结合的关键。在 MeCP2 的 NID 中引入 RTT 突变 R306C 会降低与 LEDGF 的相互作用。我们的数据显示,由于结合位点重叠,MeCP2 和 LEDGF 的多聚化相互抑制。与这一观察结果相一致的是,LEDGF的缺失导致NIH3T3细胞中的MeCP2和DNA病灶增大,这表明MeCP2-LEDGF复合物在染色质组织中发挥作用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Structure
生物-生化与分子生物学
CiteScore
8.90
自引率
1.80%
发文量
155
审稿时长
3-8 weeks
期刊介绍:
Structure aims to publish papers of exceptional interest in the field of structural biology. The journal strives to be essential reading for structural biologists, as well as biologists and biochemists that are interested in macromolecular structure and function. Structure strongly encourages the submission of manuscripts that present structural and molecular insights into biological function and mechanism. Other reports that address fundamental questions in structural biology, such as structure-based examinations of protein evolution, folding, and/or design, will also be considered. We will consider the application of any method, experimental or computational, at high or low resolution, to conduct structural investigations, as long as the method is appropriate for the biological, functional, and mechanistic question(s) being addressed. Likewise, reports describing single-molecule analysis of biological mechanisms are welcome.
In general, the editors encourage submission of experimental structural studies that are enriched by an analysis of structure-activity relationships and will not consider studies that solely report structural information unless the structure or analysis is of exceptional and broad interest. Studies reporting only homology models, de novo models, or molecular dynamics simulations are also discouraged unless the models are informed by or validated by novel experimental data; rationalization of a large body of existing experimental evidence and making testable predictions based on a model or simulation is often not considered sufficient.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


