Convergence and divergence of diploid and tetraploid cotton genomes
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
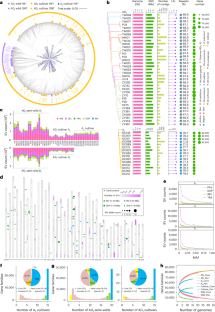
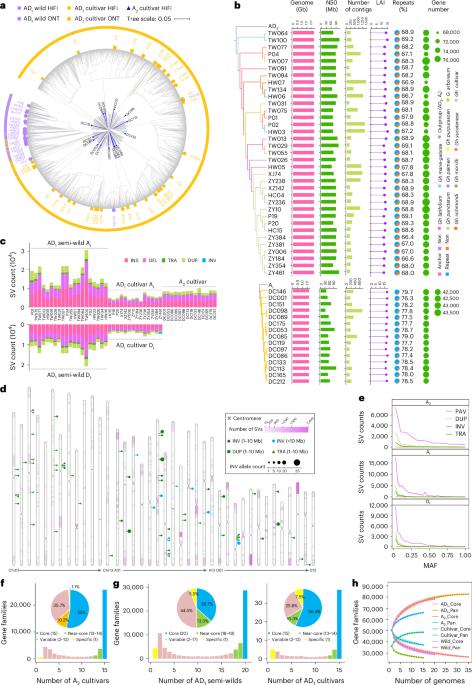
Polyploidy is an important driving force in speciation and evolution; however, the genomic basis for parallel selection of a particular trait between polyploids and ancestral diploids remains unexplored. Here we construct graph-based pan-genomes for diploid (A2) and allotetraploid (AD1) cotton species, enabled by an assembly of 50 genomes of genetically diverse accessions. We delineate a mosaic genome map of tetraploid cultivars that illustrates genomic contributions from semi-wild forms into modern cultivars. Pan-genome comparisons identify syntenic and hyper-divergent regions of continued variation between diploid and tetraploid cottons, and suggest an ongoing process of sequence evolution potentially linked to the contrasting genome size change in two subgenomes. We highlight 43% of genetic regulatory relationships for gene expression in diploid encompassing sequence divergence after polyploidy, and specifically characterize six underexplored convergent genetic loci contributing to parallel selection of fiber quality. This study offers a framework for pan-genomic dissection of genetic regulatory components underlying parallel selection of desirable traits in organisms. High-quality assemblies of 15 diploid and 35 allotetraploid cotton accessions are analyzed in graph-based pan-genomes, providing insights into genome dynamics and regulatory control of fiber transcriptomes under varying ploidy and selection pressures.

二倍体和四倍体棉花基因组的趋同与分化
多倍体是物种分化和进化的重要驱动力;然而,多倍体和祖先二倍体之间平行选择特定性状的基因组基础仍未得到探索。在这里,我们通过组装 50 个基因不同的棉花品种基因组,为二倍体(A2)和异源四倍体(AD1)棉花品种构建了基于图谱的泛基因组。我们绘制了四倍体栽培品种的马赛克基因组图谱,说明了从半野生形式到现代栽培品种的基因组贡献。泛基因组比较确定了二倍体棉花和四倍体棉花之间持续变异的同源区和超分化区,并表明序列进化的持续过程可能与两个亚基因组中基因组大小的对比变化有关。我们强调了 43% 的二倍体基因表达遗传调控关系,其中包括多倍体后的序列差异,并具体描述了六个未被充分探索的趋同遗传位点对纤维质量平行选择的贡献。这项研究为从泛基因组学角度剖析生物体内平行选择理想性状的遗传调控成分提供了一个框架。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


