Metatranscriptomics-guided discovery and characterization of a polyphenol-metabolizing gut microbial enzyme
IF 20.6
1区 医学
Q1 MICROBIOLOGY
引用次数: 0
Abstract
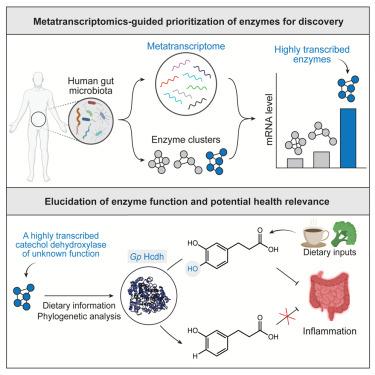
Gut microbial catechol dehydroxylases are a largely uncharacterized family of metalloenzymes that potentially impact human health by metabolizing dietary polyphenols. Here, we use metatranscriptomics (MTX) to identify highly transcribed catechol-dehydroxylase-encoding genes in human gut microbiomes. We discover a prevalent, previously uncharacterized catechol dehydroxylase (Gp Hcdh) from Gordonibacter pamelaeae that dehydroxylates hydrocaffeic acid (HCA), an anti-inflammatory gut microbial metabolite derived from plant-based foods. Further analyses suggest that the activity of Gp Hcdh may reduce anti-inflammatory benefits of polyphenol-rich foods. Together, these results show the utility of combining MTX analysis and biochemical characterization for gut microbial enzyme discovery and reveal a potential link between host inflammation and a specific polyphenol-metabolizing gut microbial enzyme.
元转录组学指导下的多酚代谢肠道微生物酶的发现与表征
肠道微生物儿茶酚脱羟化酶是一个基本未定性的金属酶家族,它通过代谢膳食中的多酚而对人体健康产生潜在影响。在这里,我们使用元转录组学(MTX)来识别人类肠道微生物组中高度转录的儿茶酚脱羟酶编码基因。我们发现了一种普遍存在的、以前未定性的儿茶酚脱羟基酶(Gp Hcdh),它来自于 Gordonibacter pamelaeae,能脱羟基化氢咖啡酸(HCA),这是一种从植物性食物中提取的抗炎性肠道微生物代谢物。进一步的分析表明,Gp Hcdh 的活性可能会降低富含多酚食物的抗炎功效。总之,这些结果表明了将 MTX 分析和生化表征结合起来发现肠道微生物酶的实用性,并揭示了宿主炎症与特定多酚代谢肠道微生物酶之间的潜在联系。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Cell host & microbe
生物-微生物学
CiteScore
45.10
自引率
1.70%
发文量
201
审稿时长
4-8 weeks
期刊介绍:
Cell Host & Microbe is a scientific journal that was launched in March 2007. The journal aims to provide a platform for scientists to exchange ideas and concepts related to the study of microbes and their interaction with host organisms at a molecular, cellular, and immune level. It publishes novel findings on a wide range of microorganisms including bacteria, fungi, parasites, and viruses. The journal focuses on the interface between the microbe and its host, whether the host is a vertebrate, invertebrate, or plant, and whether the microbe is pathogenic, non-pathogenic, or commensal. The integrated study of microbes and their interactions with each other, their host, and the cellular environment they inhabit is a unifying theme of the journal. The published work in Cell Host & Microbe is expected to be of exceptional significance within its field and also of interest to researchers in other areas. In addition to primary research articles, the journal features expert analysis, commentary, and reviews on current topics of interest in the field.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


