Post-translationally created hybrids
IF 42.8
1区 化学
Q1 CHEMISTRY, PHYSICAL
引用次数: 0
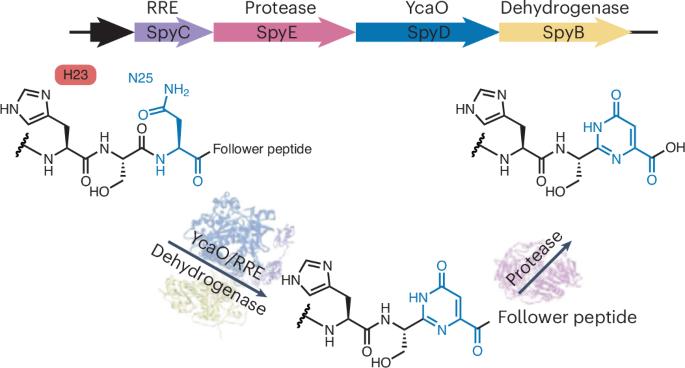
翻译后产生的杂交种
研究人员试图了解最近通过生物信息学方法发现的 RiPP 生物合成基因簇(BGC)中的蛋白质的生物合成作用。为此,在大肠杆菌中异源表达了来自链霉菌 NRRL F-5065 的同源 Spy BGC,包括其前体肽和编码蛋白 SpyC、SpyE、SpyD 和 SpyB。研究人员发现,前体酶促转化产生的肽的质量损失为 20 Da,这与丝氨酸或苏氨酸残基上形成的唑分子一致,但令人惊讶的是,进一步的实验并未证实这一点。为了便于表征,研究人员制作了该肽的缩短版,这使他们能够确定前体的修饰发生在天冬酰胺残基上。此外,核磁共振光谱显示形成了类似尿嘧啶的六元嘧啶酮环结构。根据生化分析,研究人员提出了一个分两步的途径(如图):YcaO/RRE/脱氢酶复合物(由 SpyC、SpyD 和 SpyB 组成)在底物组氨酸残基的协助下,将肽中无反应的天冬酰胺残基转化为嘧啶酮环。随后,C 端从肽被蛋白酶(SpyE)裂解,生成最终的肽-核糖核酸杂交产物。这项工作中报告的杂交分子可能会促使未来的研究解决因其发现而产生的问题。虽然参与核碱基形成的酶已经确定,但可能存在不同的催化机制。此外,杂交产物的生物功能尚不清楚。最后,这种途径在生物工程中的潜力,以及这种肽-核碱基杂交的生物合成逻辑在自然界中的广泛应用还有待探索。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Catalysis
Chemical Engineering-Bioengineering
CiteScore
52.10
自引率
1.10%
发文量
140
期刊介绍:
Nature Catalysis serves as a platform for researchers across chemistry and related fields, focusing on homogeneous catalysis, heterogeneous catalysis, and biocatalysts, encompassing both fundamental and applied studies. With a particular emphasis on advancing sustainable industries and processes, the journal provides comprehensive coverage of catalysis research, appealing to scientists, engineers, and researchers in academia and industry.
Maintaining the high standards of the Nature brand, Nature Catalysis boasts a dedicated team of professional editors, rigorous peer-review processes, and swift publication times, ensuring editorial independence and quality. The journal publishes work spanning heterogeneous catalysis, homogeneous catalysis, and biocatalysis, covering areas such as catalytic synthesis, mechanisms, characterization, computational studies, nanoparticle catalysis, electrocatalysis, photocatalysis, environmental catalysis, asymmetric catalysis, and various forms of organocatalysis.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


