Machine-guided design of cell-type-targeting cis-regulatory elements
IF 48.5
1区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 0
Abstract
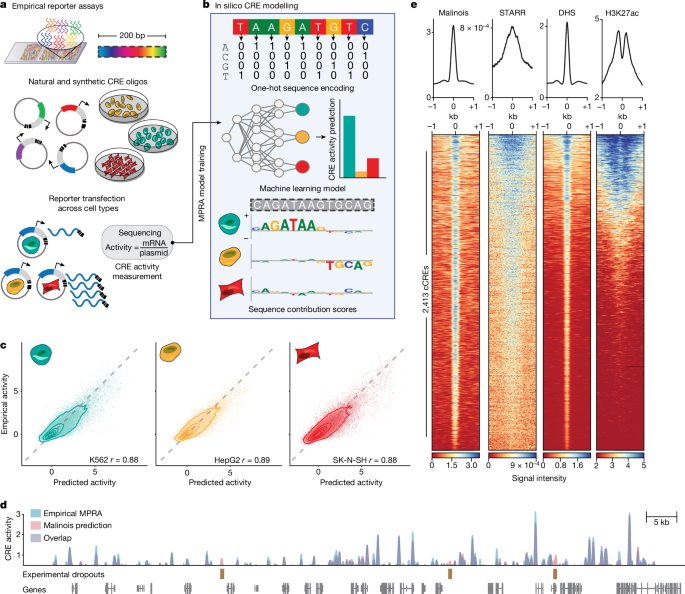
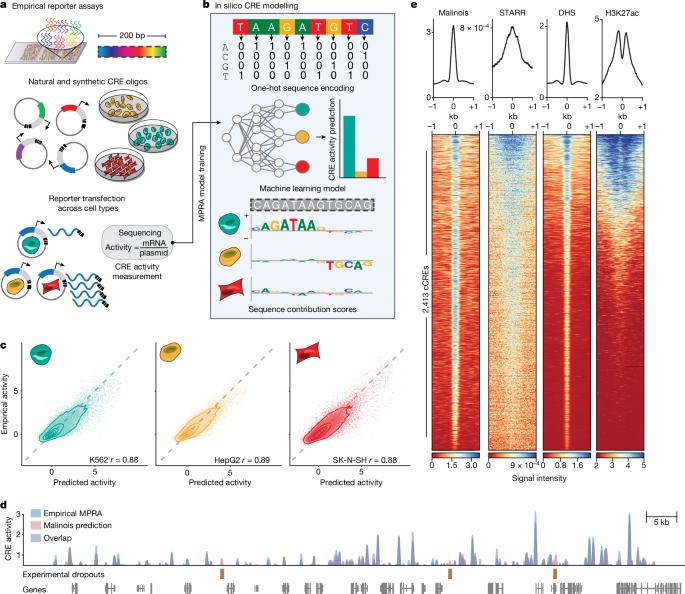
Cis-regulatory elements (CREs) control gene expression, orchestrating tissue identity, developmental timing and stimulus responses, which collectively define the thousands of unique cell types in the body1–3. While there is great potential for strategically incorporating CREs in therapeutic or biotechnology applications that require tissue specificity, there is no guarantee that an optimal CRE for these intended purposes has arisen naturally. Here we present a platform to engineer and validate synthetic CREs capable of driving gene expression with programmed cell-type specificity. We take advantage of innovations in deep neural network modelling of CRE activity across three cell types, efficient in silico optimization and massively parallel reporter assays to design and empirically test thousands of CREs4–8. Through large-scale in vitro validation, we show that synthetic sequences are more effective at driving cell-type-specific expression in three cell lines compared with natural sequences from the human genome and achieve specificity in analogous tissues when tested in vivo. Synthetic sequences exhibit distinct motif vocabulary associated with activity in the on-target cell type and a simultaneous reduction in the activity of off-target cells. Together, we provide a generalizable framework to prospectively engineer CREs from massively parallel reporter assay models and demonstrate the required literacy to write fit-for-purpose regulatory code. A generalizable framework to prospectively engineer cis-regulatory elements from massively parallel reporter assay models can be used to write fit-for-purpose regulatory code.
机器引导设计细胞类型靶向顺式调控元件
顺式调控元件(CRE)控制着基因表达,协调着组织特性、发育时间和刺激反应,共同决定着人体内成千上万种独特的细胞类型1,2,3。虽然在需要组织特异性的治疗或生物技术应用中战略性地加入 CREs 有很大的潜力,但并不能保证这些预期目的的最佳 CRE 是自然产生的。在这里,我们提出了一个平台,用于设计和验证能够以程序化细胞类型特异性驱动基因表达的合成 CRE。我们利用对三种细胞类型的 CRE 活性进行深度神经网络建模、高效硅学优化和大规模并行报告检测等方面的创新,设计并实证测试了数千种 CRE4,5,6,7,8。通过大规模体外验证,我们发现与来自人类基因组的天然序列相比,合成序列能更有效地在三种细胞系中驱动细胞特异性表达,并且在体内测试时能在类似组织中实现特异性表达。合成序列表现出与在目标细胞类型中的活性相关的不同主题词汇,同时降低了在非目标细胞中的活性。总之,我们提供了一个可通用的框架,从大规模并行报告分析模型中前瞻性地设计 CREs,并展示了编写适合目的调控代码所需的素养。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature
综合性期刊-综合性期刊
CiteScore
90.00
自引率
1.20%
发文量
3652
审稿时长
3 months
期刊介绍:
Nature is a prestigious international journal that publishes peer-reviewed research in various scientific and technological fields. The selection of articles is based on criteria such as originality, importance, interdisciplinary relevance, timeliness, accessibility, elegance, and surprising conclusions. In addition to showcasing significant scientific advances, Nature delivers rapid, authoritative, insightful news, and interpretation of current and upcoming trends impacting science, scientists, and the broader public. The journal serves a dual purpose: firstly, to promptly share noteworthy scientific advances and foster discussions among scientists, and secondly, to ensure the swift dissemination of scientific results globally, emphasizing their significance for knowledge, culture, and daily life.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


