Interpretable spatially aware dimension reduction of spatial transcriptomics with STAMP
IF 36.1
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
Abstract
Spatial transcriptomics produces high-dimensional gene expression measurements with spatial context. Obtaining a biologically meaningful low-dimensional representation of such data is crucial for effective interpretation and downstream analysis. Here, we present Spatial Transcriptomics Analysis with topic Modeling to uncover spatial Patterns (STAMP), an interpretable spatially aware dimension reduction method built on a deep generative model that returns biologically relevant, low-dimensional spatial topics and associated gene modules. STAMP can analyze data ranging from a single section to multiple sections and from different technologies to time-series data, returning topics matching known biological domains and associated gene modules containing established markers highly ranked within. In a lung cancer sample, STAMP delineated cell states with supporting markers at a higher resolution than the original annotation and uncovered cancer-associated fibroblasts concentrated on the tumor edge’s exterior. In time-series data of mouse embryonic development, STAMP disentangled the erythro-myeloid hematopoiesis and hepatocytes developmental trajectories within the liver. STAMP is highly scalable and can handle more than 500,000 cells. Spatial Transcriptomics Analysis with topic Modeling to uncover spatial Patterns (STAMP) is a versatile and scalable method for dimension reduction in spatially resolved transcriptomics that enables discovery of biologically relevant tissue domains.
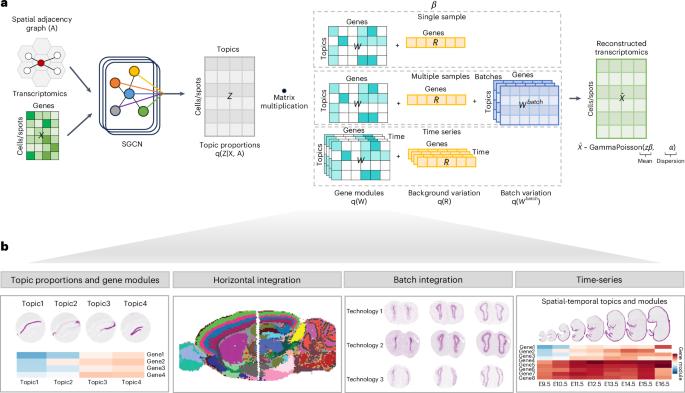
利用 STAMP 对空间转录组学进行可解释的空间感知降维。
空间转录组学产生了具有空间背景的高维基因表达测量数据。为这类数据获取具有生物学意义的低维表示对于有效解释和下游分析至关重要。在这里,我们介绍了利用主题建模揭示空间模式的空间转录组学分析(STAMP),这是一种可解释的空间感知降维方法,它建立在一个深度生成模型之上,可返回生物相关的低维空间主题和相关基因模块。STAMP 可以分析从单个切片到多个切片的数据,以及从不同技术到时间序列的数据,并返回与已知生物领域相匹配的主题和相关的基因模块,这些基因模块中包含已建立的高度排序的标记物。在肺癌样本中,STAMP 以比原始注释更高的分辨率划分了带有支持标记的细胞状态,并发现了集中在肿瘤边缘外部的癌症相关成纤维细胞。在小鼠胚胎发育的时间序列数据中,STAMP 分离了肝脏内红细胞-髓系造血和肝细胞的发育轨迹。STAMP 具有高度可扩展性,可处理超过 500,000 个细胞。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Methods
生物-生化研究方法
CiteScore
58.70
自引率
1.70%
发文量
326
审稿时长
1 months
期刊介绍:
Nature Methods is a monthly journal that focuses on publishing innovative methods and substantial enhancements to fundamental life sciences research techniques. Geared towards a diverse, interdisciplinary readership of researchers in academia and industry engaged in laboratory work, the journal offers new tools for research and emphasizes the immediate practical significance of the featured work. It publishes primary research papers and reviews recent technical and methodological advancements, with a particular interest in primary methods papers relevant to the biological and biomedical sciences. This includes methods rooted in chemistry with practical applications for studying biological problems.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


