Systematic discovery of antibacterial and antifungal bacterial toxins
IF 19.4
1区 生物学
Q1 MICROBIOLOGY
引用次数: 0
Abstract
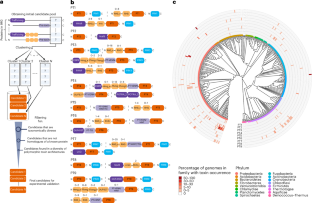
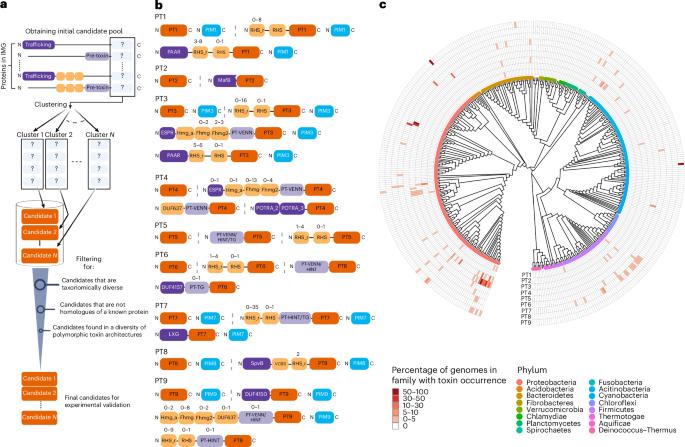
Microorganisms use toxins to kill competing microorganisms or eukaryotic cells. Polymorphic toxins are proteins that encode carboxy-terminal toxin domains. Here we developed a computational approach to identify previously undiscovered, conserved toxin domains of polymorphic toxins within 105,438 microbial genomes. We validated nine short toxins, showing that they cause cell death upon heterologous expression in either Escherichia coli or Saccharomyces cerevisiae. Five cognate immunity genes that neutralize the toxins were also discovered. The toxins are encoded by 2.2% of sequenced bacteria. A subset of the toxins exhibited potent antifungal activity against various pathogenic fungi but not against two invertebrate model organisms or macrophages. Experimental validation suggested that these toxins probably target the cell membrane or DNA or inhibit cell division. Further characterization and structural analysis of two toxin–immunity protein complexes confirmed DNase activity. These findings expand our knowledge of microbial toxins involved in inter-microbial competition that may have the potential for clinical and biotechnological applications. Genome sequence mining and computational analyses lead to the discovery and functional characterization of conserved bacterial toxins with activity against bacteria and fungi.
抗菌和抗真菌细菌毒素的系统发现
微生物利用毒素杀死竞争微生物或真核细胞。多态毒素是编码羧基末端毒素结构域的蛋白质。在这里,我们开发了一种计算方法,在 105,438 个微生物基因组中识别出以前未被发现的多态毒素保守毒素结构域。我们验证了九种短毒素,发现它们在大肠杆菌或酿酒酵母中异源表达时会导致细胞死亡。我们还发现了五个能中和毒素的同源免疫基因。2.2%的测序细菌编码这些毒素。这些毒素的一个子集对多种致病真菌具有很强的抗真菌活性,但对两种无脊椎动物模式生物或巨噬细胞没有活性。实验验证表明,这些毒素可能以细胞膜或 DNA 为靶标,或抑制细胞分裂。对两种毒素-免疫蛋白复合物的进一步表征和结构分析证实了 DNase 活性。这些发现拓展了我们对参与微生物间竞争的微生物毒素的认识,可能具有临床和生物技术应用潜力。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature Microbiology
Immunology and Microbiology-Microbiology
CiteScore
44.40
自引率
1.10%
发文量
226
期刊介绍:
Nature Microbiology aims to cover a comprehensive range of topics related to microorganisms. This includes:
Evolution: The journal is interested in exploring the evolutionary aspects of microorganisms. This may include research on their genetic diversity, adaptation, and speciation over time.
Physiology and cell biology: Nature Microbiology seeks to understand the functions and characteristics of microorganisms at the cellular and physiological levels. This may involve studying their metabolism, growth patterns, and cellular processes.
Interactions: The journal focuses on the interactions microorganisms have with each other, as well as their interactions with hosts or the environment. This encompasses investigations into microbial communities, symbiotic relationships, and microbial responses to different environments.
Societal significance: Nature Microbiology recognizes the societal impact of microorganisms and welcomes studies that explore their practical applications. This may include research on microbial diseases, biotechnology, or environmental remediation.
In summary, Nature Microbiology is interested in research related to the evolution, physiology and cell biology of microorganisms, their interactions, and their societal relevance.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


