Base editing screens define the genetic landscape of cancer drug resistance mechanisms
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
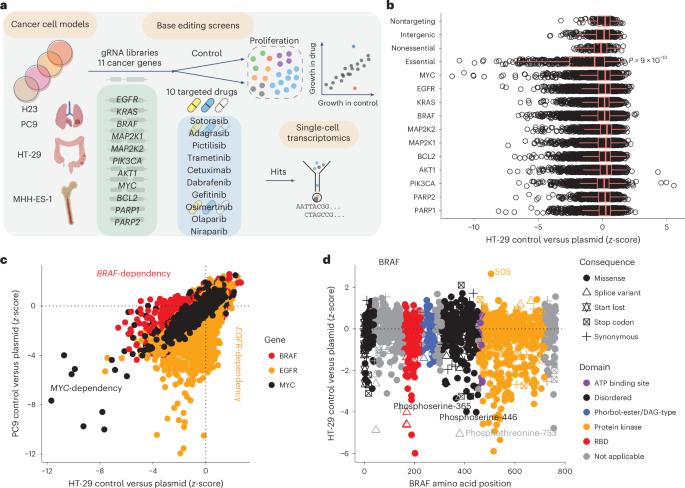
Drug resistance is a principal limitation to the long-term efficacy of cancer therapies. Cancer genome sequencing can retrospectively delineate the genetic basis of drug resistance, but this requires large numbers of post-treatment samples to nominate causal variants. Here we prospectively identify genetic mechanisms of resistance to ten oncology drugs from CRISPR base editing mutagenesis screens in four cancer cell lines using a guide RNA library predicted to install 32,476 variants in 11 cancer genes. We identify four functional classes of protein variants modulating drug sensitivity and use single-cell transcriptomics to reveal how these variants operate through distinct mechanisms, including eliciting a drug-addicted cell state. We identify variants that can be targeted with alternative inhibitors to overcome resistance and functionally validate an epidermal growth factor receptor (EGFR) variant that sensitizes lung cancer cells to EGFR inhibitors. Our variant-to-function map has implications for patient stratification, therapy combinations and drug scheduling in cancer treatment. Base editing screens of 11 cancer genes identify four functional classes of variants that collectively underpin sensitivity and resistance to ten commonly used drugs in cancer cell lines.

碱基编辑筛选确定癌症耐药机制的基因图谱
耐药性是癌症疗法长期疗效的主要限制因素。癌症基因组测序可以回顾性地描述耐药性的遗传基础,但这需要大量的治疗后样本来确定因果变异。在这里,我们通过 CRISPR 碱基编辑诱变筛选,在四种癌细胞系中使用引导 RNA 文库,预测在 11 个癌症基因中安装了 32,476 个变体,从而前瞻性地确定了十种肿瘤药物耐药性的遗传机制。我们确定了调节药物敏感性的四类功能性蛋白质变体,并利用单细胞转录组学揭示了这些变体如何通过不同的机制发挥作用,包括诱发药物成瘾的细胞状态。我们发现了可以用替代抑制剂克服耐药性的变体,并从功能上验证了表皮生长因子受体(EGFR)变体能使肺癌细胞对 EGFR 抑制剂敏感。我们的变体-功能图谱对癌症治疗中的患者分层、治疗组合和药物调配具有重要意义。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


