Mimicking a Cellular Crowding Environment for Enzyme-Free Paper-Based Nucleic Acid Tests at the Point of Care
IF 8.2
1区 化学
Q1 CHEMISTRY, ANALYTICAL
引用次数: 0
Abstract
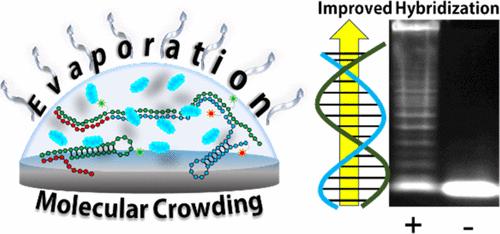
Point of care (PoC) nucleic acid amplification tests (NAATs) are a cornerstone of public health, providing the earliest and most accurate diagnostic method for many communicable diseases in the same location where the patient receives treatment. Communicable diseases, such as human immunodeficiency virus (HIV), disproportionately impact low-resource communities where NAATs are often unobtainable due to the resource-intensive enzymes that drive the tests. Enzyme-free nucleic acid detection methods, such as hybridization chain reaction (HCR), use DNA secondary structures for self-driven amplification schemes, producing large DNA nanostructures, capable of single-molecule detection in cellulo. These thermodynamically driven DNA-based tests have struggled to penetrate the PoC diagnostic field due to their inadequate limits of detection or complex workflows. Here, we present a proof-of-concept NAAT that combines HCR-based amplification of a target nucleic acid sequence with paper-based nucleic acid filtration and enrichment capable of detecting sub-pM levels of synthetic DNA. We reconstruct the favorable hybridization conditions of an in cellulo reaction in vitro by incubating HCR in an evaporating, microvolume environment containing poly(ethylene glycol) as a crowding agent. We demonstrate that the kinetics and thermodynamics of DNA–DNA and DNA–RNA hybridization is enhanced by the dynamic evaporating environment and inclusion of crowding agents, bringing HCR closer to meeting PoC NAAT needs.
在医疗点模拟细胞拥挤环境进行无酶纸基核酸检测
医疗点核酸扩增检测(NAATs)是公共卫生的基石,可在患者接受治疗的同一地点为许多传染病提供最早、最准确的诊断方法。人类免疫缺陷病毒(HIV)等传染病对资源匮乏的社区造成了极大的影响,而在这些社区,由于需要使用耗费大量资源的酶来进行检测,因此往往无法获得 NAAT。无酶核酸检测方法,如杂交链反应(HCR),利用 DNA 二级结构进行自驱动扩增,产生大型 DNA 纳米结构,能够在细胞内进行单分子检测。这些基于热力学驱动的 DNA 检测方法由于检测极限不足或工作流程复杂,一直难以渗透到 PoC 诊断领域。在这里,我们介绍一种概念验证型 NAAT,它将基于 HCR 的目标核酸序列扩增与基于纸的核酸过滤和富集相结合,能够检测亚 pM 水平的合成 DNA。我们通过在含有聚乙二醇(poly(ethylene glycol)作为拥挤剂的蒸发微体积环境中培养 HCR,重建了体外细胞内反应的有利杂交条件。我们证明,DNA-DNA 和 DNA-RNA 杂交的动力学和热力学在动态蒸发环境和加入拥挤剂后得到增强,从而使 HCR 更接近于满足 PoC NAAT 的需求。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

ACS Sensors
Chemical Engineering-Bioengineering
CiteScore
14.50
自引率
3.40%
发文量
372
期刊介绍:
ACS Sensors is a peer-reviewed research journal that focuses on the dissemination of new and original knowledge in the field of sensor science, particularly those that selectively sense chemical or biological species or processes. The journal covers a broad range of topics, including but not limited to biosensors, chemical sensors, gas sensors, intracellular sensors, single molecule sensors, cell chips, and microfluidic devices. It aims to publish articles that address conceptual advances in sensing technology applicable to various types of analytes or application papers that report on the use of existing sensing concepts in new ways or for new analytes.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


