Saturation mutagenesis-reinforced functional assays for disease-related genes
IF 5.7
1区 化学
Q2 CHEMISTRY, PHYSICAL
引用次数: 0
Abstract
Interpretation of disease-causing genetic variants remains a challenge in human genetics. Current costs and complexity of deep mutational scanning methods are obstacles for achieving genome-wide resolution of variants in disease-related genes. Our framework, saturation mutagenesis-reinforced functional assays (SMuRF), offers simple and cost-effective saturation mutagenesis paired with streamlined functional assays to enhance the interpretation of unresolved variants. Applying SMuRF to neuromuscular disease genes FKRP and LARGE1, we generated functional scores for all possible coding single-nucleotide variants, which aid in resolving clinically reported variants of uncertain significance. SMuRF also demonstrates utility in predicting disease severity, resolving critical structural regions, and providing training datasets for the development of computational predictors. Overall, our approach enables variant-to-function insights for disease genes in a cost-effective manner that can be broadly implemented by standard research laboratories.
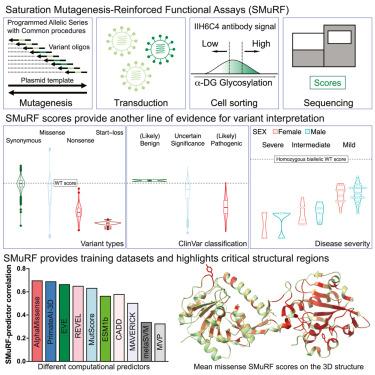
疾病相关基因的饱和诱变强化功能测试
解读致病基因变异仍是人类遗传学的一项挑战。目前深度突变扫描方法的成本和复杂性是实现疾病相关基因变异全基因组解析的障碍。我们的框架--饱和诱变-强化功能测定(SMuRF)--提供了简单、经济的饱和诱变方法,并配以简化的功能测定,以加强对未解决变异的解释。将 SMuRF 应用于神经肌肉疾病基因 FKRP 和 LARGE1,我们为所有可能的编码单核苷酸变异生成了功能评分,这有助于解决临床报告的意义不确定的变异。SMuRF 在预测疾病严重程度、确定关键结构区域以及为开发计算预测器提供训练数据集方面也显示出了实用性。总之,我们的方法能以经济高效的方式深入了解疾病基因的变异到功能,可在标准研究实验室广泛应用。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Journal of Chemical Theory and Computation
化学-物理:原子、分子和化学物理
CiteScore
9.90
自引率
16.40%
发文量
568
审稿时长
1 months
期刊介绍:
The Journal of Chemical Theory and Computation invites new and original contributions with the understanding that, if accepted, they will not be published elsewhere. Papers reporting new theories, methodology, and/or important applications in quantum electronic structure, molecular dynamics, and statistical mechanics are appropriate for submission to this Journal. Specific topics include advances in or applications of ab initio quantum mechanics, density functional theory, design and properties of new materials, surface science, Monte Carlo simulations, solvation models, QM/MM calculations, biomolecular structure prediction, and molecular dynamics in the broadest sense including gas-phase dynamics, ab initio dynamics, biomolecular dynamics, and protein folding. The Journal does not consider papers that are straightforward applications of known methods including DFT and molecular dynamics. The Journal favors submissions that include advances in theory or methodology with applications to compelling problems.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


