Integration of variant annotations using deep set networks boosts rare variant association testing
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
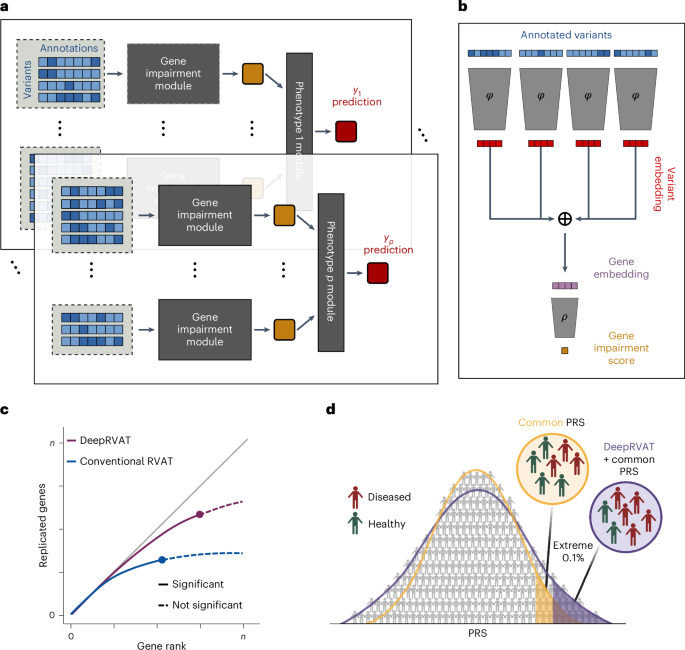
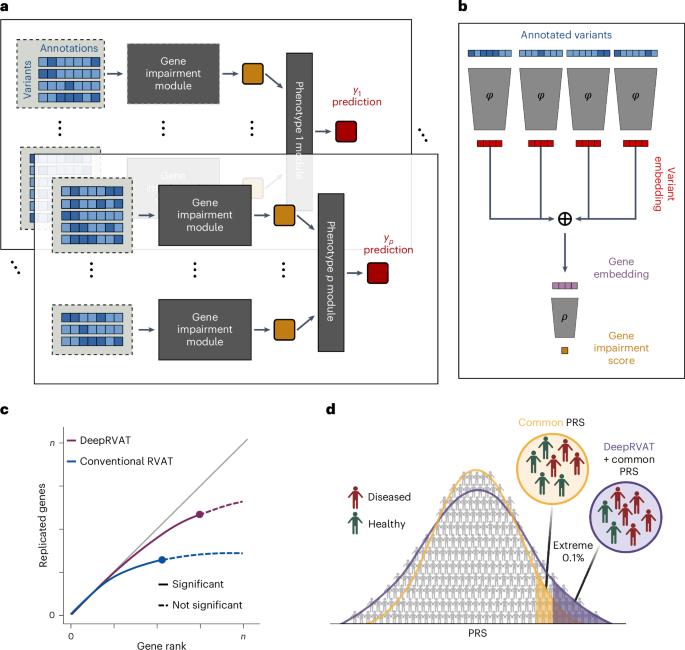
Rare genetic variants can have strong effects on phenotypes, yet accounting for rare variants in genetic analyses is statistically challenging due to the limited number of allele carriers and the burden of multiple testing. While rich variant annotations promise to enable well-powered rare variant association tests, methods integrating variant annotations in a data-driven manner are lacking. Here we propose deep rare variant association testing (DeepRVAT), a model based on set neural networks that learns a trait-agnostic gene impairment score from rare variant annotations and phenotypes, enabling both gene discovery and trait prediction. On 34 quantitative and 63 binary traits, using whole-exome-sequencing data from UK Biobank, we find that DeepRVAT yields substantial gains in gene discoveries and improved detection of individuals at high genetic risk. Finally, we demonstrate how DeepRVAT enables calibrated and computationally efficient rare variant tests at biobank scale, aiding the discovery of genetic risk factors for human disease traits. Deep rare variant association testing (DeepRVAT) is a deep set neural network model that flexibly integrates rare variant annotations into a trait-agnostic gene impairment score. These scores improve association testing and polygenic risk prediction.
利用深度集网络整合变异注释可促进罕见变异关联测试
罕见遗传变异对表型有很大影响,但由于等位基因携带者数量有限和多重测试的负担,在遗传分析中考虑罕见变异在统计学上具有挑战性。虽然丰富的变异注释有望实现强大的罕见变异关联测试,但目前还缺乏以数据驱动方式整合变异注释的方法。在此,我们提出了深度罕见变异关联测试(DeepRVAT),这是一种基于集合神经网络的模型,它能从罕见变异注释和表型中学习与性状无关的基因损伤评分,从而实现基因发现和性状预测。我们利用英国生物库的全外显子测序数据对 34 个定量性状和 63 个二元性状进行了分析,发现 DeepRVAT 在发现基因和检测高遗传风险个体方面都有显著提高。最后,我们展示了 DeepRVAT 如何在生物库规模上实现校准和计算高效的罕见变异测试,从而帮助发现人类疾病特征的遗传风险因素。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


