Single-cell and bulk transcriptional profiling of mouse ovaries reveals novel genes and pathways associated with DNA damage response in oocytes
IF 2.1
3区 生物学
Q2 DEVELOPMENTAL BIOLOGY
引用次数: 0
Abstract
Immature oocytes enclosed in primordial follicles stored in female ovaries are under constant threat of DNA damage induced by endogenous and exogenous factors. Checkpoint kinase 2 (CHEK2) is a key mediator of the DNA damage response (DDR) in all cells. Genetic studies have shown that CHEK2 and its downstream targets, p53, and TAp63, regulate primordial follicle elimination in response to DNA damage. However, the mechanism leading to their demise is still poorly characterized. Single-cell and bulk RNA sequencing were used to determine the DDR in wild-type and Chek2-deficient ovaries. A low but oocyte-lethal dose of ionizing radiation induces ovarian DDR that is solely dependent on CHEK2. DNA damage activates multiple response pathways related to apoptosis, p53, interferon signaling, inflammation, cell adhesion, and intercellular communication. These pathways are differentially employed by different ovarian cell types, with oocytes disproportionately affected by radiation. Novel genes and pathways are induced by radiation specifically in oocytes, shedding light on their sensitivity to DNA damage, and implicating a coordinated response between oocytes and pregranulosa cells within the follicle. These findings provide a foundation for future studies on the specific mechanisms regulating oocyte survival in the context of aging, therapeutic and environmental genotoxic exposures.
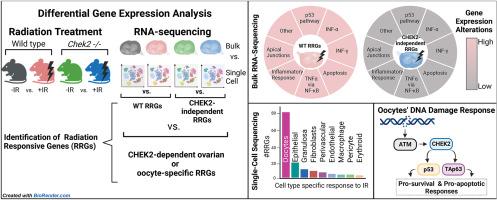
小鼠卵巢单细胞和大容量转录谱分析揭示了与卵母细胞 DNA 损伤反应相关的新型基因和通路。
储存在女性卵巢原始卵泡中的未成熟卵母细胞不断受到内源性和外源性因素诱导的 DNA 损伤的威胁。检查点激酶2(CHEK2)是所有细胞中DNA损伤反应(DDR)的关键介质。遗传学研究表明,CHEK2 及其下游靶标 p53 和 TAp63 在 DNA 损伤反应中调节原始卵泡的消亡。然而,导致原始卵泡消亡的机制仍不甚明了。研究人员利用单细胞和大量RNA测序来确定野生型和Chek2缺陷型卵巢的DDR。低剂量但卵母细胞致死的电离辐射诱导卵巢DDR完全依赖于CHEK2。DNA 损伤会激活与细胞凋亡、p53、干扰素信号、炎症、细胞粘附和细胞间通讯有关的多种反应途径。不同类型的卵巢细胞采用不同的途径,其中卵母细胞受辐射的影响尤为严重。新的基因和途径在卵母细胞中受到辐射的特异性诱导,揭示了卵母细胞对DNA损伤的敏感性,并暗示了卵母细胞和卵泡内前颗粒细胞之间的协调反应。这些发现为今后研究在衰老、治疗和环境基因毒性暴露的背景下调节卵母细胞存活的具体机制奠定了基础。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Developmental biology
生物-发育生物学
CiteScore
5.30
自引率
3.70%
发文量
182
审稿时长
1.5 months
期刊介绍:
Developmental Biology (DB) publishes original research on mechanisms of development, differentiation, and growth in animals and plants at the molecular, cellular, genetic and evolutionary levels. Areas of particular emphasis include transcriptional control mechanisms, embryonic patterning, cell-cell interactions, growth factors and signal transduction, and regulatory hierarchies in developing plants and animals.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


