Influence of Software Settings on the Identification Rate, Quantification Results, and Reproducibility in Profiling Post-Translational Modifications by Microflow Liquid Chromatography–Ion Mobility-Quadrupole Time-Of-Flight Analysis Using PEAKS Software
IF 3.8
2区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
Abstract
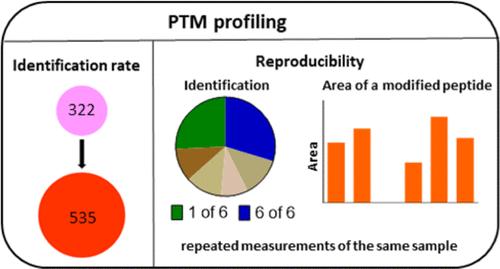
The influence of data evaluation parameters on qualitative and quantitative results of untargeted shotgun profiling of enzymatic and nonenzymatic post-translational modifications (PTMs) was investigated in a model of bovine whey protein α-lactalbumin heated with lactose. Based on the same raw data, individual adjustments to the protein database and enzyme settings of PEAKS studio software increased the identification rate from 27 unmodified peptides to 48 and from 322 peptides in total to 535. The qualitative and quantitative reproducibility was also assessed based on 18 measurements of one sample across three batches. A total of 570 peptides were detected. While 89 peptides were identified in all measurements, the majority of peptides (161) were detected only once and mostly based on nonindicative spectra. The reproducibility of label-free quantification (LFQ) in six measurements of the same sample was similar after processing the data by either the PTM algorithm or the LFQ algorithm. In both cases, about one-third of the peptides showed a coefficient of variation of above 20%. However, the LFQ algorithm increased the number of quantified peptides from 75 to 179. Data are available at the PRIDE Archive with the data set identifier PXD050363.
使用 PEAKS 软件进行微流液相色谱-离子迁移率-四极杆飞行时间分析时,软件设置对翻译后修饰的识别率、定量结果和重现性的影响
在用乳糖加热牛乳清蛋白α-乳白蛋白的模型中,研究了数据评估参数对酶和非酶翻译后修饰(PTMs)非靶向猎枪谱分析定性和定量结果的影响。基于相同的原始数据,对蛋白质数据库和 PEAKS studio 软件的酶设置进行了个别调整,使识别率从 27 个未修饰肽段提高到 48 个,肽段总数从 322 个提高到 535 个。此外,还根据一个样品在三个批次中的 18 次测量结果,对定性和定量的可重复性进行了评估。共检测到 570 个肽段。虽然在所有测量中都鉴定出了 89 个肽段,但大多数肽段(161 个)只被检测到一次,而且大多是基于非指示性光谱。在使用 PTM 算法或 LFQ 算法处理数据后,对同一样品进行的六次测量中,无标记定量(LFQ)的重现性相似。在这两种情况下,都有约三分之一的肽变异系数超过 20%。不过,LFQ 算法将量化肽段的数量从 75 个增加到 179 个。数据可在 PRIDE Archive 上查阅,数据集标识符为 PXD050363。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Journal of Proteome Research
生物-生化研究方法
CiteScore
9.00
自引率
4.50%
发文量
251
审稿时长
3 months
期刊介绍:
Journal of Proteome Research publishes content encompassing all aspects of global protein analysis and function, including the dynamic aspects of genomics, spatio-temporal proteomics, metabonomics and metabolomics, clinical and agricultural proteomics, as well as advances in methodology including bioinformatics. The theme and emphasis is on a multidisciplinary approach to the life sciences through the synergy between the different types of "omics".
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


