Small Regulatory RNAs of the Rsm Clan in Pseudomonas
IF 2.6
2区 生物学
Q3 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
Bacteria of the genus Pseudomonas are ubiquitous on Earth due to their great metabolic versatility and adaptation to fluctuating environments and different hosts. Some groups are important animal/human and plant pathogens, whereas others are studied for their biotechnological applications, including bioremediation, biological control of phytopathogens and plant growth promotion. Notably, their adaptability is mediated by various signal transduction systems, with the post-transcriptional Gac-Rsm cascade playing a key role. This pervasive Pseudomonas pathway controls major transitions at the population level, such as motile/sessile lifestyle, primary/secondary metabolism or replicative/infective behaviour. A hallmark of the Gac-Rsm cascade is the participation of small, regulatory, non-coding RNAs of the Rsm clan. These RNAs are synthetised in response to cell-density-dependent autoinducer signals channelled through the GacS/GacA two-component system, and they counteract, by molecular mimicry, the translational control that RNA-binding proteins of the RsmA family exert over hundreds of mRNAs. Rsm RNAs have been investigated in a few Pseudomonas model species, evidencing the presence of a variable number and families of genes depending on the taxonomic clade. However, the global picture of the distribution of these riboregulators at the genus level was unknown until now. We have undertaken a comprehensive survey and annotation of the vast array of gene sequences encoding members of the Rsm RNA clan in 245 complete genomes that cover 28 phylogenomic clades across the entire genus. The properties of the different families of rsm genes, their phylogenetic radiation, as well as the features of their promoters and adjacent regions, are discussed. The novel insights presented in our manuscript will significantly boost research on the biology of these prevalent RNAs in understudied species of the genus Pseudomonas and closely related genera.
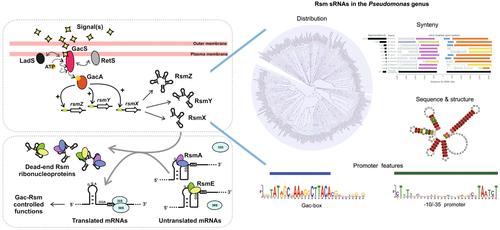
假单胞菌 Rsm 家族的小调控 RNA
假单胞菌属细菌在地球上无处不在,这是因为它们的代谢能力很强,能适应多变的环境和不同的宿主。一些假单胞菌属是重要的动物/人类和植物病原体,而另一些则因其生物技术应用而受到研究,包括生物修复、植物病原体生物防治和植物生长促进。值得注意的是,它们的适应性是由各种信号转导系统介导的,其中转录后 Gac-Rsm 级联起着关键作用。假单胞菌的这一普遍途径控制着种群水平上的重大转变,如运动/无运动生活方式、初级/次级新陈代谢或复制/感染行为。Gac-Rsm 级联的一个特点是 Rsm 家族的小型、调控性、非编码 RNA 的参与。这些 RNA 通过 GacS/GacA 双组分系统对依赖于细胞密度的自诱导信号做出反应而合成,它们通过分子模拟抵消了 RsmA 家族的 RNA 结合蛋白对数百种 mRNA 的翻译控制。对一些假单胞菌模式物种中的 Rsm RNA 进行了研究,结果表明,根据分类学支系的不同,存在着不同数量和家族的基因。然而,直到现在,这些核调控因子在属一级的整体分布情况仍不为人知。我们对 245 个完整基因组中编码 Rsm RNA 家族成员的大量基因序列进行了全面的调查和注释,这些基因组涵盖了全属 28 个系统发生学支系。我们讨论了不同 rsm 基因家族的特性、它们的系统发育辐射以及启动子和邻近区域的特征。我们手稿中提出的新见解将极大地推动对假单胞菌属及其近缘属中未被充分研究的物种中这些普遍存在的 RNA 的生物学研究。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Molecular Microbiology
生物-生化与分子生物学
CiteScore
7.20
自引率
5.60%
发文量
132
审稿时长
1.7 months
期刊介绍:
Molecular Microbiology, the leading primary journal in the microbial sciences, publishes molecular studies of Bacteria, Archaea, eukaryotic microorganisms, and their viruses.
Research papers should lead to a deeper understanding of the molecular principles underlying basic physiological processes or mechanisms. Appropriate topics include gene expression and regulation, pathogenicity and virulence, physiology and metabolism, synthesis of macromolecules (proteins, nucleic acids, lipids, polysaccharides, etc), cell biology and subcellular organization, membrane biogenesis and function, traffic and transport, cell-cell communication and signalling pathways, evolution and gene transfer. Articles focused on host responses (cellular or immunological) to pathogens or on microbial ecology should be directed to our sister journals Cellular Microbiology and Environmental Microbiology, respectively.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


