Proteomic Characterization of Transfusable Blood Components: Fresh Frozen Plasma, Cryoprecipitate, and Derived Extracellular Vesicles via Data-Independent Mass Spectrometry
IF 3.6
2区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
Abstract
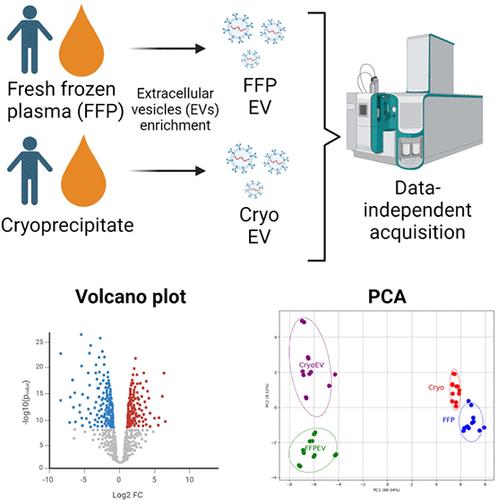
Extracellular vesicles (EVs) are a heterogeneous collection of particles that play a crucial role in cell-to-cell communication, primarily due to their ability to transport molecules, such as proteins. Thus, profiling EV-associated proteins offers insight into their biological effects. EVs can be isolated from various biological fluids, including donor blood components such as cryoprecipitate and fresh frozen plasma (FFP). In this study, we conducted a proteomic analysis of five single donor units of cryoprecipitate, FFP, and EVs derived from these blood components using a quantitative mass spectrometry approach. EVs were successfully isolated from both cryoprecipitate and FFP based on community guidelines. We identified and quantified approximately 360 proteins across all sample groups. Principal component analysis and heatmaps revealed that both cryoprecipitate and FFP are similar. Similarly, EVs derived from cryoprecipitate and FFP are comparable. However, they differ between the originating fluids and their derived EVs. Using the R-package MS-DAP, differentially expressed proteins (DEPs) were identified. The DEPs for all comparisons, when submitted for gene enrichment analysis, are involved in the complement and coagulation pathways. The protein profile generated from this study will have important clinical implications in increasing our knowledge of the proteins that are associated with EVs derived from blood components.
可输血成分的蛋白质组特征:通过数据独立质谱法鉴定新鲜冷冻血浆、冷沉淀和衍生细胞外小泡
细胞外囊泡(EVs)是一种异质的颗粒集合,在细胞间通信中发挥着至关重要的作用,这主要是由于它们具有运输蛋白质等分子的能力。因此,分析 EV 相关蛋白有助于深入了解它们的生物效应。EV可从各种生物液体中分离出来,包括低温沉淀物和新鲜冰冻血浆(FFP)等供体血液成分。在这项研究中,我们采用定量质谱方法对五个单一供体单位的低温沉淀物、新鲜冰冻血浆和从这些血液成分中提取的 EVs 进行了蛋白质组学分析。根据社区指南,我们成功地从冷冻沉淀和 FFP 中分离出了 EVs。我们鉴定并量化了所有样本组中的约 360 种蛋白质。主成分分析和热图显示,冷冻沉淀和 FFP 具有相似性。同样,从冷冻沉淀和 FFP 中提取的 EV 也具有可比性。然而,它们在原液和衍生 EV 之间存在差异。使用 R 软件包 MS-DAP,鉴定了差异表达蛋白(DEPs)。在提交基因富集分析时,所有比较的 DEPs 都参与了补体和凝血途径。这项研究生成的蛋白质图谱将对临床产生重要影响,有助于我们进一步了解与血液成分中的 EVs 相关的蛋白质。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Journal of Proteome Research
生物-生化研究方法
CiteScore
9.00
自引率
4.50%
发文量
251
审稿时长
3 months
期刊介绍:
Journal of Proteome Research publishes content encompassing all aspects of global protein analysis and function, including the dynamic aspects of genomics, spatio-temporal proteomics, metabonomics and metabolomics, clinical and agricultural proteomics, as well as advances in methodology including bioinformatics. The theme and emphasis is on a multidisciplinary approach to the life sciences through the synergy between the different types of "omics".
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


