Custom microfluidic chip design enables cost-effective three-dimensional spatiotemporal transcriptomics with a wide field of view
IF 31.7
1区 生物学
Q1 GENETICS & HEREDITY
引用次数: 0
Abstract
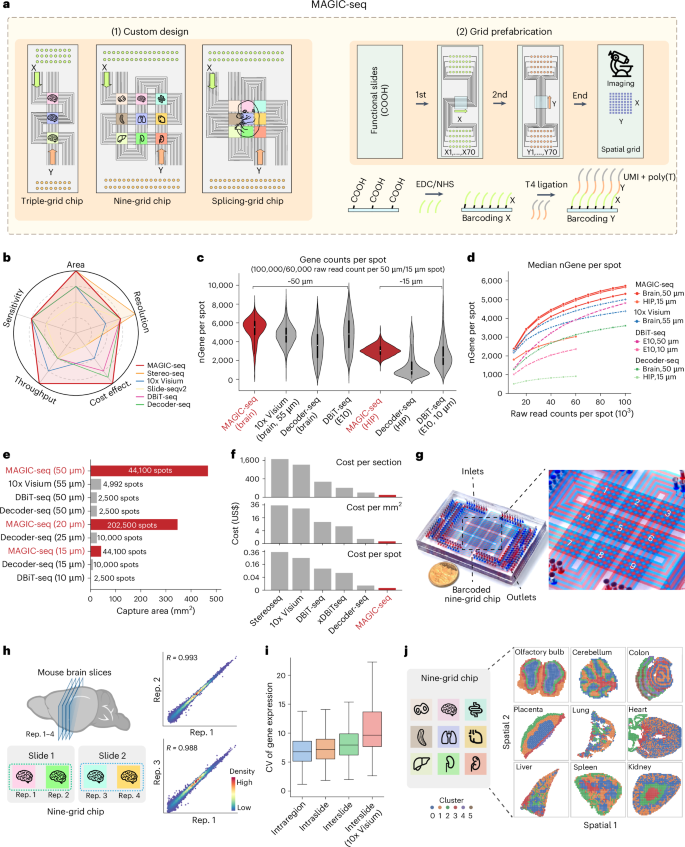
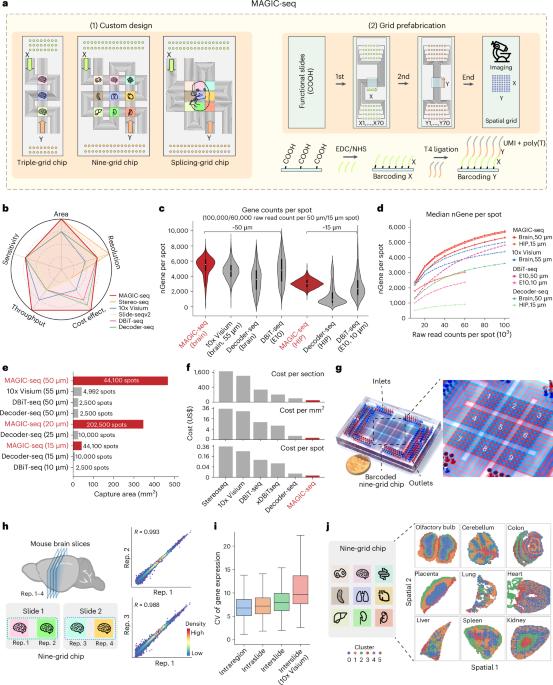
Spatial transcriptomic techniques offer unprecedented insights into the molecular organization of complex tissues. However, integrating cost-effectiveness, high throughput, a wide field of view and compatibility with three-dimensional (3D) volumes has been challenging. Here we introduce microfluidics-assisted grid chips for spatial transcriptome sequencing (MAGIC-seq), a new method that combines carbodiimide chemistry, spatial combinatorial indexing and innovative microfluidics design. This technique allows sensitive and reproducible profiling of diverse tissue types, achieving an eightfold increase in throughput, minimal cost and reduced batch effects. MAGIC-seq breaks conventional microfluidics limits by enhancing barcoding efficiency and enables analysis of whole postnatal mouse sections, providing comprehensive cellular structure elucidation at near single-cell resolution, uncovering transcriptional variations and dynamic trajectories of mouse organogenesis. Our 3D transcriptomic atlas of the developing mouse brain, consisting of 93 sections, reveals the molecular and cellular landscape, serving as a valuable resource for neuroscience and developmental biology. Overall, MAGIC-seq is a high-throughput, cost-effective, large field of view and versatile method for spatial transcriptomic studies. Microfluidics-assisted grid chips for spatial transcriptome sequencing (MAGIC-seq) is a spatial transcriptomics method combining multiple-grid microfluidic design and prefabricated DNA arrays for increased throughput and reduced cost, with applications for large fields of view and 3D spatial mapping.
定制微流控芯片设计实现了经济高效的三维时空转录组学,视野开阔
空间转录组学技术为了解复杂组织的分子组织提供了前所未有的洞察力。然而,将成本效益、高通量、宽视场以及与三维(3D)体积的兼容性融为一体一直是个挑战。在这里,我们介绍了用于空间转录组测序的微流体辅助网格芯片(MAGIC-seq),这是一种结合了碳化二亚胺化学、空间组合索引和创新微流体设计的新方法。该技术可对不同类型的组织进行灵敏且可重复的分析,将通量提高了八倍,并将成本降至最低,减少了批次效应。MAGIC-seq 打破了传统微流控技术的限制,提高了条形码效率,能够分析小鼠出生后的整个切片,以接近单细胞的分辨率提供全面的细胞结构阐释,揭示小鼠器官形成过程中的转录变化和动态轨迹。我们的发育中小鼠大脑三维转录组图谱由 93 个切片组成,揭示了分子和细胞结构,是神经科学和发育生物学的宝贵资源。总之,MAGIC-seq 是一种高通量、高性价比、大视场和多功能的空间转录组研究方法。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Nature genetics
生物-遗传学
CiteScore
43.00
自引率
2.60%
发文量
241
审稿时长
3 months
期刊介绍:
Nature Genetics publishes the very highest quality research in genetics. It encompasses genetic and functional genomic studies on human and plant traits and on other model organisms. Current emphasis is on the genetic basis for common and complex diseases and on the functional mechanism, architecture and evolution of gene networks, studied by experimental perturbation.
Integrative genetic topics comprise, but are not limited to:
-Genes in the pathology of human disease
-Molecular analysis of simple and complex genetic traits
-Cancer genetics
-Agricultural genomics
-Developmental genetics
-Regulatory variation in gene expression
-Strategies and technologies for extracting function from genomic data
-Pharmacological genomics
-Genome evolution
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


