Embryonic genome instability upon DNA replication timing program emergence
IF 50.5
1区 综合性期刊
Q1 MULTIDISCIPLINARY SCIENCES
引用次数: 0
Abstract
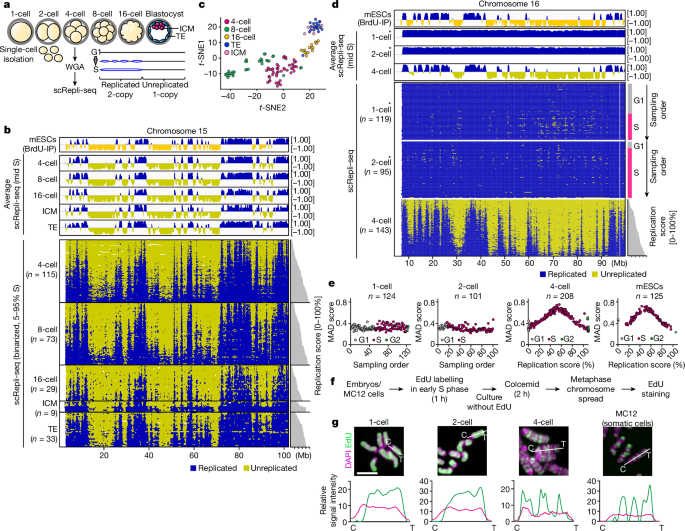
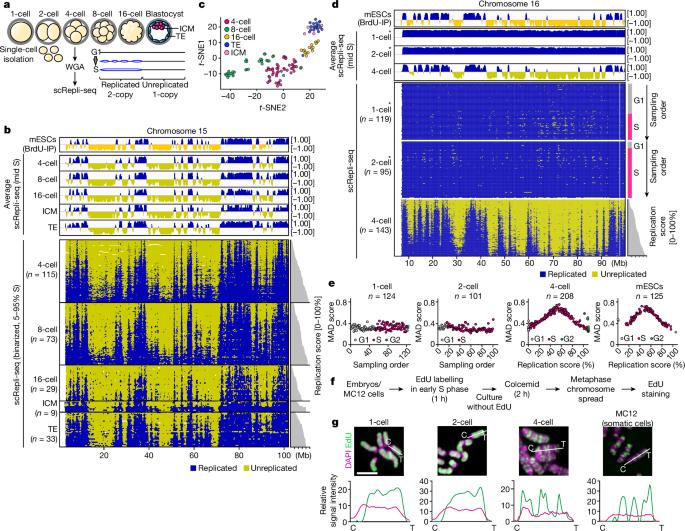
Faithful DNA replication is essential for genome integrity1–4. Under-replicated DNA leads to defects in chromosome segregation, which are common during embryogenesis5–8. However, the regulation of DNA replication remains poorly understood in early mammalian embryos. Here we constructed a single-cell genome-wide DNA replication atlas of pre-implantation mouse embryos and identified an abrupt replication program switch accompanied by a transient period of genomic instability. In 1- and 2-cell embryos, we observed the complete absence of a replication timing program, and the entire genome replicated gradually and uniformly using extremely slow-moving replication forks. In 4-cell embryos, a somatic-cell-like replication timing program commenced abruptly. However, the fork speed was still slow, S phase was extended, and markers of replication stress, DNA damage and repair increased. This was followed by an increase in break-type chromosome segregation errors specifically during the 4-to-8-cell division with breakpoints enriched in late-replicating regions. These errors were rescued by nucleoside supplementation, which accelerated fork speed and reduced the replication stress. By the 8-cell stage, forks gained speed, S phase was no longer extended and chromosome aberrations decreased. Thus, a transient period of genomic instability exists during normal mouse development, preceded by an S phase lacking coordination between replisome-level regulation and megabase-scale replication timing regulation, implicating a link between their coordination and genome stability. A single-cell genome-wide DNA replication atlas of pre-implantation mouse embryos reveals an abrupt replication program switch accompanied by a transient period of genomic instability.
DNA 复制定时程序出现时胚胎基因组的不稳定性
可靠的 DNA 复制对基因组的完整性至关重要1,2,3,4。DNA 复制不足会导致染色体分离缺陷,这在胚胎发生过程中很常见5,6,7,8。然而,人们对哺乳动物早期胚胎中 DNA 复制的调控仍然知之甚少。在这里,我们构建了植入前小鼠胚胎的单细胞全基因组 DNA 复制图谱,并确定了突然的复制程序转换伴随着短暂的基因组不稳定性。在 1 细胞和 2 细胞胚胎中,我们观察到完全没有复制定时程序,整个基因组利用移动速度极慢的复制叉逐渐均匀地复制。在 4 细胞胚胎中,类似体细胞的复制定时程序突然开始。然而,复制叉的速度仍然很慢,S 期延长,复制压力、DNA 损伤和修复的标志物增加。随后,断裂型染色体分离错误增加,特别是在 4 至 8 细胞分裂期间,断裂点集中在晚期复制区域。补充核苷可加快分叉速度并减轻复制压力,从而挽救这些错误。到了 8 细胞阶段,分叉速度加快,S 期不再延长,染色体畸变减少。因此,在小鼠正常发育过程中存在一个短暂的基因组不稳定期,在此之前的S期缺乏复制体级调控和巨酶级复制定时调控之间的协调,这意味着它们之间的协调与基因组稳定性之间存在联系。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature
综合性期刊-综合性期刊
CiteScore
90.00
自引率
1.20%
发文量
3652
审稿时长
3 months
期刊介绍:
Nature is a prestigious international journal that publishes peer-reviewed research in various scientific and technological fields. The selection of articles is based on criteria such as originality, importance, interdisciplinary relevance, timeliness, accessibility, elegance, and surprising conclusions. In addition to showcasing significant scientific advances, Nature delivers rapid, authoritative, insightful news, and interpretation of current and upcoming trends impacting science, scientists, and the broader public. The journal serves a dual purpose: firstly, to promptly share noteworthy scientific advances and foster discussions among scientists, and secondly, to ensure the swift dissemination of scientific results globally, emphasizing their significance for knowledge, culture, and daily life.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


