Efficient HLA imputation from sequential SNPs data by transformer
IF 2.6
3区 生物学
Q2 GENETICS & HEREDITY
引用次数: 0
Abstract
Human leukocyte antigen (HLA) genes are associated with a variety of diseases, yet the direct typing of HLA alleles is both time-consuming and costly. Consequently, various imputation methods leveraging sequential single nucleotide polymorphisms (SNPs) data have been proposed, employing either statistical or deep learning models, such as the convolutional neural network (CNN)-based model, DEEP*HLA. However, these methods exhibit limited imputation efficiency for infrequent alleles and necessitate a large size of reference dataset. In this context, we have developed a Transformer-based model to HLA allele imputation, named “HLA Reliable IMpuatioN by Transformer (HLARIMNT)” designed to exploit the sequential nature of SNPs data. We evaluated HLARIMNT’s performance using two distinct reference panels; Pan-Asian reference panel (n = 530) and Type 1 Diabetes genetics Consortium (T1DGC) reference panel (n = 5225), alongside a combined panel (n = 1060). HLARIMNT demonstrated superior accuracy to DEEP*HLA across several indices, particularly for infrequent alleles. Furthermore, we explored the impact of varying training data sizes on imputation accuracy, finding that HLARIMNT consistently outperformed across all data size. These findings suggest that Transformer-based models can efficiently impute not only HLA types but potentially other gene types from sequential SNPs data.
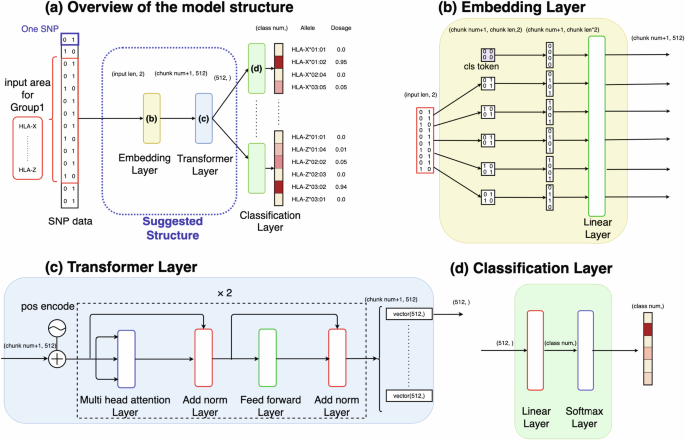
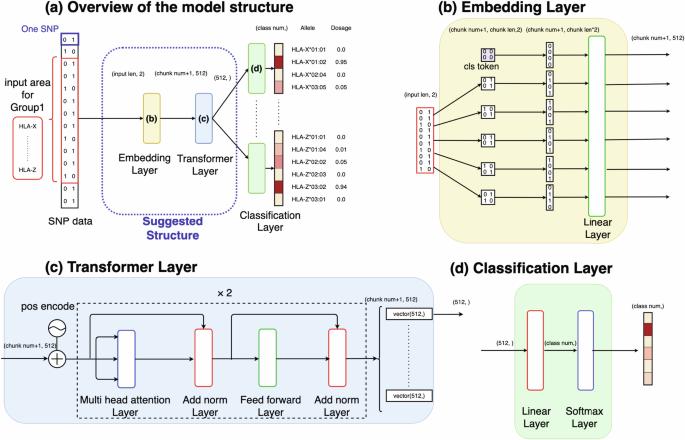
通过转换器从序列 SNPs 数据中高效推算 HLA。
人类白细胞抗原(HLA)基因与多种疾病相关,但直接进行 HLA 等位基因分型既费时又费钱。因此,人们提出了各种利用序列单核苷酸多态性(SNPs)数据的估算方法,采用统计或深度学习模型,如基于卷积神经网络(CNN)的模型 DEEP*HLA。然而,这些方法对于不常见的等位基因的估算效率有限,而且需要大量的参考数据集。在这种情况下,我们开发了一种基于变换器的 HLA 等位基因估算模型,命名为 "HLA Reliable IMpuatioN by Transformer (HLARIMNT)",旨在利用 SNPs 数据的连续性。我们使用两个不同的参考面板(泛亚参考面板(n = 530)和 1 型糖尿病遗传学联盟(T1DGC)参考面板(n = 5225))以及一个组合面板(n = 1060)评估了 HLARIMNT 的性能。在多个指标上,HLARIMNT 的准确性都优于 DEEP*HLA,特别是对于不常见的等位基因。此外,我们还探讨了不同训练数据规模对估算准确性的影响,发现在所有数据规模下,HLARIMNT 的表现始终优于 DEEP*HLA。这些研究结果表明,基于 Transformer 的模型不仅能有效地归因 HLA 类型,还可能从序列 SNPs 数据中归因其他基因类型。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Journal of Human Genetics
生物-遗传学
CiteScore
7.20
自引率
0.00%
发文量
101
审稿时长
4-8 weeks
期刊介绍:
The Journal of Human Genetics is an international journal publishing articles on human genetics, including medical genetics and human genome analysis. It covers all aspects of human genetics, including molecular genetics, clinical genetics, behavioral genetics, immunogenetics, pharmacogenomics, population genetics, functional genomics, epigenetics, genetic counseling and gene therapy.
Articles on the following areas are especially welcome: genetic factors of monogenic and complex disorders, genome-wide association studies, genetic epidemiology, cancer genetics, personal genomics, genotype-phenotype relationships and genome diversity.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


