Genome-resolved metagenomics: a game changer for microbiome medicine
IF 9.5
2区 医学
Q1 BIOCHEMISTRY & MOLECULAR BIOLOGY
引用次数: 0
Abstract
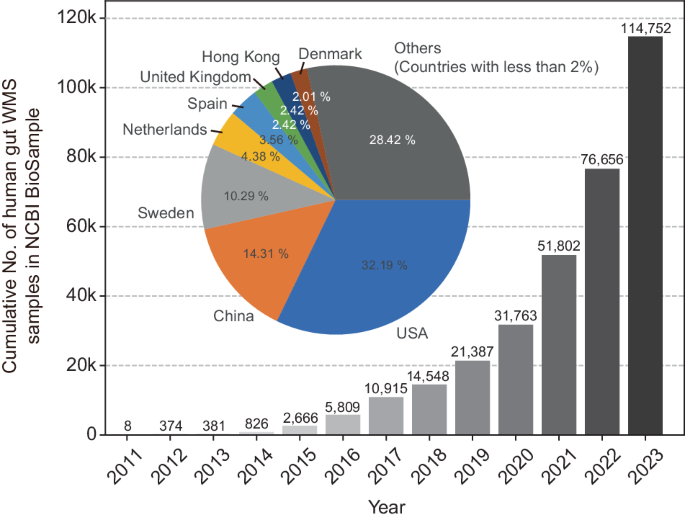
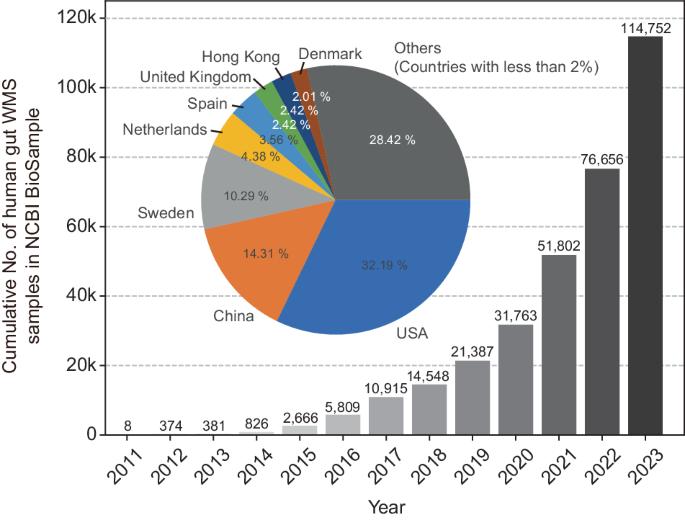
Recent substantial evidence implicating commensal bacteria in human diseases has given rise to a new domain in biomedical research: microbiome medicine. This emerging field aims to understand and leverage the human microbiota and derivative molecules for disease prevention and treatment. Despite the complex and hierarchical organization of this ecosystem, most research over the years has relied on 16S amplicon sequencing, a legacy of bacterial phylogeny and taxonomy. Although advanced sequencing technologies have enabled cost-effective analysis of entire microbiota, translating the relatively short nucleotide information into the functional and taxonomic organization of the microbiome has posed challenges until recently. In the last decade, genome-resolved metagenomics, which aims to reconstruct microbial genomes directly from whole-metagenome sequencing data, has made significant strides and continues to unveil the mysteries of various human-associated microbial communities. There has been a rapid increase in the volume of whole metagenome sequencing data and in the compilation of novel metagenome-assembled genomes and protein sequences in public depositories. This review provides an overview of the capabilities and methods of genome-resolved metagenomics for studying the human microbiome, with a focus on investigating the prokaryotic microbiota of the human gut. Just as decoding the human genome and its variations marked the beginning of the genomic medicine era, unraveling the genomes of commensal microbes and their sequence variations is ushering us into the era of microbiome medicine. Genome-resolved metagenomics stands as a pivotal tool in this transition and can accelerate our journey toward achieving these scientific and medical milestones. The human body houses numerous microbes, tiny organisms, that are vital for our health. This research aims to overcome limitations using genome-resolved metagenomics, a method that assembles complete genomes from complex microbial communities without needing to grow the organisms in a lab. The study focuses on the gut microbiome, using advanced computer methods to build metagenome-assembled genomes from DNA sequencing data. The research successfully increased the genetic diversity of the human gut microbiome by adding many new genomes to the existing database. The main findings include identifying new microbial species and expanding the genetic repertoire of known species, providing deeper understanding of the microbial diversity within the human gut. Researchers conclude that genome-resolved metagenomics is a significant advancement in microbiome research, offering understanding of microbial communities and their functions. This summary was initially drafted using artificial intelligence, then revised and fact-checked by the author.
基因组解析元基因组学:改变微生物组医学的游戏规则。
最近有大量证据表明,共生细菌与人类疾病有关联,这催生了生物医学研究的一个新领域:微生物组医学。这一新兴领域旨在了解和利用人体微生物群及其衍生分子来预防和治疗疾病。尽管这一生态系统的组织结构复杂且层次分明,但多年来大多数研究都依赖于 16S 扩增子测序,这是细菌系统发育和分类学的遗产。尽管先进的测序技术能够对整个微生物群进行经济有效的分析,但直到最近,将相对较短的核苷酸信息转化为微生物群的功能和分类组织仍是一项挑战。近十年来,旨在直接从全基因组测序数据重建微生物基因组的基因组解析元基因组学取得了长足的进步,并不断揭开各种人类相关微生物群落的神秘面纱。全元基因组测序数据的数量以及在公共储存库中汇编的新型元基因组组装基因组和蛋白质序列都在迅速增加。本综述概述了基因组分辨元基因组学研究人类微生物组的能力和方法,重点是研究人类肠道的原核微生物群。正如解码人类基因组及其变异标志着基因组医学时代的开始一样,揭示共生微生物的基因组及其序列变异正把我们带入微生物组医学时代。基因组解析元基因组学是这一转变过程中的关键工具,可以加快我们实现这些科学和医学里程碑的进程。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊

Experimental and Molecular Medicine
医学-生化与分子生物学
CiteScore
19.50
自引率
0.80%
发文量
166
审稿时长
3 months
期刊介绍:
Experimental & Molecular Medicine (EMM) stands as Korea's pioneering biochemistry journal, established in 1964 and rejuvenated in 1996 as an Open Access, fully peer-reviewed international journal. Dedicated to advancing translational research and showcasing recent breakthroughs in the biomedical realm, EMM invites submissions encompassing genetic, molecular, and cellular studies of human physiology and diseases. Emphasizing the correlation between experimental and translational research and enhanced clinical benefits, the journal actively encourages contributions employing specific molecular tools. Welcoming studies that bridge basic discoveries with clinical relevance, alongside articles demonstrating clear in vivo significance and novelty, Experimental & Molecular Medicine proudly serves as an open-access, online-only repository of cutting-edge medical research.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


