Morphological and molecular preservation through universal preparation of fresh-frozen tissue samples for multimodal imaging workflows
IF 13.1
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
Abstract
The landscape of tissue-based imaging modalities is constantly and rapidly evolving. While formalin-fixed, paraffin-embedded material is still useful for histological imaging, the fixation process irreversibly changes the molecular composition of the sample. Therefore, many imaging approaches require fresh-frozen material to get meaningful results. This is particularly true for molecular imaging techniques such as mass spectrometry imaging, which are widely used to probe the spatial arrangement of the tissue metabolome. As high-quality fresh-frozen tissues are limited in their availability, any sample preparation workflow they are subjected to needs to ensure morphological and molecular preservation of the tissues and be compatible with as many of the established and emerging imaging techniques as possible to obtain the maximum possible insights from the tissues. Here we describe a universal sample preparation workflow, from the initial step of freezing the tissues to the cold embedding in a new hydroxypropyl methylcellulose/polyvinylpyrrolidone-enriched hydrogel and the generation of thin tissue sections for analysis. Moreover, we highlight the optimized storage conditions that limit molecular and morphological degradation of the sections. The protocol is compatible with human and plant tissues and can be easily adapted for the preparation of alternative sample formats (e.g., three-dimensional cell cultures). The integrated workflow is universally compatible with histological tissue analysis, mass spectrometry imaging and imaging mass cytometry, as well as spatial proteomic, genomic and transcriptomic tissue analysis. The protocol can be completed within 4 h and requires minimal prior experience in the preparation of tissue samples for multimodal imaging experiments. The morphological and molecular preservation of fresh-frozen tissues is difficult. Embedding with an hydroxypropyl methylcellulose/polyvinylpyrrolidone-rich hydrogel results in a material compatible with spatial biochemical analysis (e.g., mass spectrometry imaging), enabling multimodal data integration.
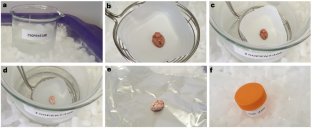
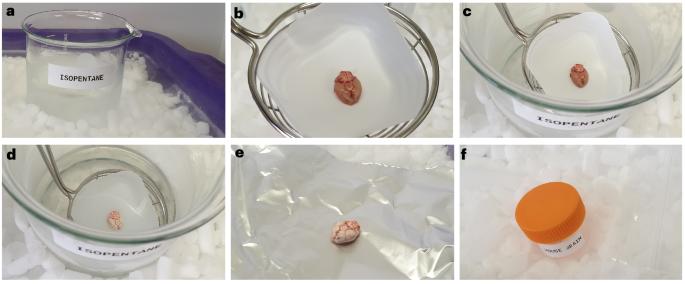
通过为多模态成像工作流程普遍制备新鲜冷冻组织样本,实现形态和分子保存。
基于组织的成像模式在不断快速发展。虽然福尔马林固定、石蜡包埋材料仍可用于组织学成像,但固定过程会不可逆地改变样本的分子组成。因此,许多成像方法需要新鲜冷冻材料才能获得有意义的结果。质谱成像等分子成像技术尤其如此,这些技术被广泛用于探测组织代谢组的空间排列。由于高质量的新鲜冷冻组织供应有限,因此任何样本制备工作流程都必须确保组织的形态和分子保存,并尽可能与多种成熟和新兴成像技术兼容,以便从组织中获得最大可能的洞察力。在此,我们介绍一种通用的样本制备工作流程,从冷冻组织的初始步骤,到冷包埋在新型羟丙基甲基纤维素/聚乙烯吡咯烷酮富含的水凝胶中,再到生成用于分析的薄组织切片。此外,我们还强调了限制切片分子和形态降解的优化储存条件。该方案与人体组织和植物组织兼容,并可轻松适用于制备其他形式的样品(如三维细胞培养物)。集成工作流程与组织学组织分析、质谱成像、成像质谱以及空间蛋白质组、基因组和转录组组织分析普遍兼容。该方案可在 4 小时内完成,只需具备为多模态成像实验制备组织样本的最低经验。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature Protocols
生物-生化研究方法
CiteScore
29.10
自引率
0.70%
发文量
128
审稿时长
4 months
期刊介绍:
Nature Protocols focuses on publishing protocols used to address significant biological and biomedical science research questions, including methods grounded in physics and chemistry with practical applications to biological problems. The journal caters to a primary audience of research scientists and, as such, exclusively publishes protocols with research applications. Protocols primarily aimed at influencing patient management and treatment decisions are not featured.
The specific techniques covered encompass a wide range, including but not limited to: Biochemistry, Cell biology, Cell culture, Chemical modification, Computational biology, Developmental biology, Epigenomics, Genetic analysis, Genetic modification, Genomics, Imaging, Immunology, Isolation, purification, and separation, Lipidomics, Metabolomics, Microbiology, Model organisms, Nanotechnology, Neuroscience, Nucleic-acid-based molecular biology, Pharmacology, Plant biology, Protein analysis, Proteomics, Spectroscopy, Structural biology, Synthetic chemistry, Tissue culture, Toxicology, and Virology.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


