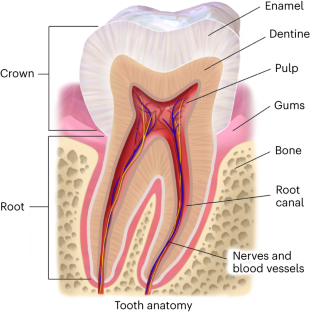
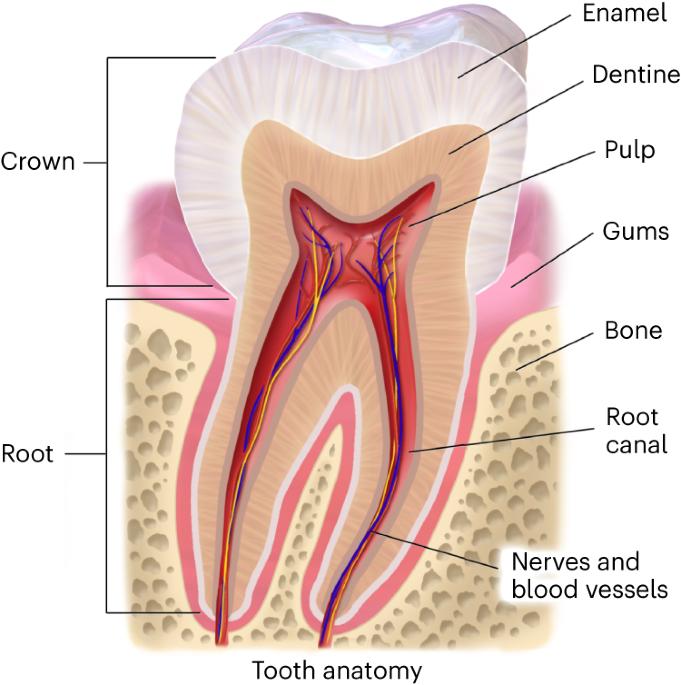
Deep-time phylogenetic inference by paleoproteomic analysis of dental enamel
IF 13.1
1区 生物学
Q1 BIOCHEMICAL RESEARCH METHODS
引用次数: 0
Abstract
In temperate and subtropical regions, ancient proteins are reported to survive up to about 2 million years, far beyond the known limits of ancient DNA preservation in the same areas. Accordingly, their amino acid sequences currently represent the only source of genetic information available to pursue phylogenetic inference involving species that went extinct too long ago to be amenable for ancient DNA analysis. Here we present a complete workflow, including sample preparation, mass spectrometric data acquisition and computational analysis, to recover and interpret million-year-old dental enamel protein sequences. During sample preparation, the proteolytic digestion step, usually an integral part of conventional bottom-up proteomics, is omitted to increase the recovery of the randomly degraded peptides spontaneously generated by extensive diagenetic hydrolysis of ancient proteins over geological time. Similarly, we describe other solutions we have adopted to (1) authenticate the endogenous origin of the protein traces we identify, (2) detect and validate amino acid variation in the ancient protein sequences and (3) attempt phylogenetic inference. Sample preparation and data acquisition can be completed in 3–4 working days, while subsequent data analysis usually takes 2–5 days. The workflow described requires basic expertise in ancient biomolecules analysis, mass spectrometry-based proteomics and molecular phylogeny. Finally, we describe the limits of this approach and its potential for the reconstruction of evolutionary relationships in paleontology and paleoanthropology. Ancient proteins carry genetic information from fossils that are too old or degraded for ancient DNA recovery. This protocol describes the extraction and tandem mass spectrometry sequencing of million-year-old dental enamel proteins for phylogenetic inference.
通过牙釉质古蛋白组分析进行深时系统发育推断
据报道,在温带和亚热带地区,古蛋白质的存活时间长达约 200 万年,远远超出了已知的古 DNA 在同一地区的保存极限。因此,它们的氨基酸序列是目前唯一可用来进行系统发育推断的遗传信息来源,涉及的物种在很久以前就已经灭绝,不适合进行古 DNA 分析。在这里,我们介绍了一个完整的工作流程,包括样品制备、质谱数据采集和计算分析,以恢复和解释百万年前的牙釉质蛋白质序列。在样品制备过程中,我们省略了通常是传统自下而上蛋白质组学不可或缺的一部分的蛋白质分解步骤,以提高对古蛋白质在地质年代中大量成因水解自发产生的随机降解肽的回收率。同样,我们还介绍了我们采用的其他解决方案,以便:(1) 鉴定蛋白质踪迹的内源来源;(2) 检测和验证古蛋白质序列中的氨基酸变异;(3) 尝试进行系统发育推断。样品制备和数据采集可在 3-4 个工作日内完成,而后续的数据分析通常需要 2-5 天。所述工作流程要求具备古代生物分子分析、基于质谱的蛋白质组学和分子系统学方面的基本专业知识。最后,我们将介绍这种方法的局限性及其在重建古生物学和古人类学进化关系方面的潜力。
本文章由计算机程序翻译,如有差异,请以英文原文为准。
求助全文
约1分钟内获得全文
求助全文
来源期刊
Nature Protocols
生物-生化研究方法
CiteScore
29.10
自引率
0.70%
发文量
128
审稿时长
4 months
期刊介绍:
Nature Protocols focuses on publishing protocols used to address significant biological and biomedical science research questions, including methods grounded in physics and chemistry with practical applications to biological problems. The journal caters to a primary audience of research scientists and, as such, exclusively publishes protocols with research applications. Protocols primarily aimed at influencing patient management and treatment decisions are not featured.
The specific techniques covered encompass a wide range, including but not limited to: Biochemistry, Cell biology, Cell culture, Chemical modification, Computational biology, Developmental biology, Epigenomics, Genetic analysis, Genetic modification, Genomics, Imaging, Immunology, Isolation, purification, and separation, Lipidomics, Metabolomics, Microbiology, Model organisms, Nanotechnology, Neuroscience, Nucleic-acid-based molecular biology, Pharmacology, Plant biology, Protein analysis, Proteomics, Spectroscopy, Structural biology, Synthetic chemistry, Tissue culture, Toxicology, and Virology.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


