基于FCBF法和叠加集成模型的硝化位点预测
IF 0.5
4区 生物学
Q4 BIOCHEMICAL RESEARCH METHODS
引用次数: 3
摘要
硝化作用是发生在蛋白质酪氨酸残基上的重要翻译后修饰(PTM)之一。蛋白酪氨酸硝化在疾病条件下的发生是不可避免的,它代表了从no信号转导生理作用向氧化和潜在致病途径的转变。异常的蛋白质硝化修饰可导致严重的人类疾病,包括神经退行性疾病、急性呼吸窘迫、器官移植排斥反应和肺癌。确定蛋白质序列中的硝化位点是十分必要和重要的。预测蛋白质序列中哪些酪氨酸残基被硝化,哪些不被硝化,对研究硝化机理和相关疾病具有重要意义。在本研究中,提出了一种基于过欠采样策略和FCBF方法的硝化位点预测模型,通过叠加集成学习和融合多个特征。首先,用2701维融合特征(PseAAC、PSSM、aindex、CKSAAP、Disorder)对蛋白序列样本进行编码。其次,根据对称不确定性度量,采用FCBF方法生成排序特征集;第三,在模型训练过程中,采用过采样和欠采样技术处理不平衡数据集。最后,采用增量特征选择(IFS)方法提取基于10次交叉验证的最优分类器。结果表明,无论在独立测试集上与其他分类器进行比较,还是在训练集上与单一类型特征或融合特征进行交叉验证,该模型在MCC、Recall、F1-score等指标上都具有显著的性能优势。通过综合FCBF特征排序方法、过采样和欠采样技术以及多基分类器组成的叠加模型,建立了有效的硝化PTM位点预测模型,该模型在正负样本比例高度不平衡的情况下能够达到较高的召回率。本文章由计算机程序翻译,如有差异,请以英文原文为准。
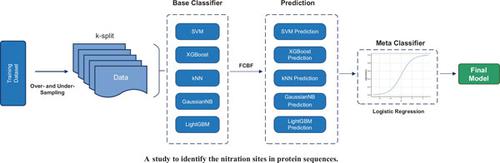
Prediction of Nitration Sites Based on FCBF Method and Stacking Ensemble Model
Nitration is one of the important Post-Translational Modification (PTM) occurring on the tyrosine residues of proteins. The occurrence of protein tyrosine nitration under disease conditions is inevitable and represents a shift from the signal transducing physiological actions of -NO to oxidative and potentially pathogenic pathways. Abnormal protein nitration modification can lead to serious human diseases, including neurodegenerative diseases, acute respiratory distress, organ transplant rejection and lung cancer. It is necessary and important to identify the nitration sites in protein sequences. Predicting that which tyrosine residues in the protein sequence are nitrated and which are not is of great significance for the study of nitration mechanism and related diseases. In this study, a prediction model of nitration sites based on the over-under sampling strategy and the FCBF method was proposed by stacking ensemble learning and fusing multiple features. Firstly, the protein sequence sample was encoded by 2701-dimensional fusion features (PseAAC, PSSM, AAIndex, CKSAAP, Disorder). Secondly, the ranked feature set was generated by the FCBF method according to the symmetric uncertainty metric. Thirdly, in the process of model training, use the over- and under- sampling technique was used to tackle the imbalanced dataset. Finally, the Incremental Feature Selection (IFS) method was adopted to extract an optimal classifier based on 10-fold cross-validation. Results show that the model has significant performance advantages in indicators such as MCC, Recall and F1-score, no matter in what way the comparison was conducted with other classifiers on the independent test set, or made by cross-validation with single-type feature or with fusion-features on the training set. By integrating the FCBF feature ranking methods, over- and under- sampling technique and a stacking model composed of multiple base classifiers, an effective prediction model for nitration PTM sites was build, which can achieve a better recall rate when the ratio of positive and negative samples is highly imbalanced.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Current Proteomics
BIOCHEMICAL RESEARCH METHODS-BIOCHEMISTRY & MOLECULAR BIOLOGY
CiteScore
1.60
自引率
0.00%
发文量
25
审稿时长
>0 weeks
期刊介绍:
Research in the emerging field of proteomics is growing at an extremely rapid rate. The principal aim of Current Proteomics is to publish well-timed in-depth/mini review articles in this fast-expanding area on topics relevant and significant to the development of proteomics. Current Proteomics is an essential journal for everyone involved in proteomics and related fields in both academia and industry.
Current Proteomics publishes in-depth/mini review articles in all aspects of the fast-expanding field of proteomics. All areas of proteomics are covered together with the methodology, software, databases, technological advances and applications of proteomics, including functional proteomics. Diverse technologies covered include but are not limited to:
Protein separation and characterization techniques
2-D gel electrophoresis and image analysis
Techniques for protein expression profiling including mass spectrometry-based methods and algorithms for correlative database searching
Determination of co-translational and post- translational modification of proteins
Protein/peptide microarrays
Biomolecular interaction analysis
Analysis of protein complexes
Yeast two-hybrid projects
Protein-protein interaction (protein interactome) pathways and cell signaling networks
Systems biology
Proteome informatics (bioinformatics)
Knowledge integration and management tools
High-throughput protein structural studies (using mass spectrometry, nuclear magnetic resonance and X-ray crystallography)
High-throughput computational methods for protein 3-D structure as well as function determination
Robotics, nanotechnology, and microfluidics.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


