前瞻性快速全基因组测序与美国危重儿科患者匹配回顾性队列的真实世界经济评估
IF 2.9
3区 医学
Q2 GENETICS & HEREDITY
引用次数: 6
摘要
通过证明快速全基因组测序(rWGS)的实际价值来支持其应用的需求日益增长。我们旨在评估 rWGS 在病因不明的儿科重症患者中的成本效益。我们前瞻性地收集了2018年3月至2020年9月期间尼克劳斯儿童医院重症监护病房收治的rWGS患者的数据(N = 65)。在一个匹配的回顾性队列中收集了标准诊断基因检测的比较数据。我们确定了总成本、诊断收益 (DY) 和增量成本效益比 (ICER),并对选择偏差和右删减进行了调整。rWGS 为 39.8% 的患者带来了诊断结果,而标准检测仅为 13.5% (p = 0.026)。rWGS 为每人平均节省了 100,440 美元(SE = 26,497, p <0.001),为 65 名患者总共节省了 653 万美元。本文章由计算机程序翻译,如有差异,请以英文原文为准。
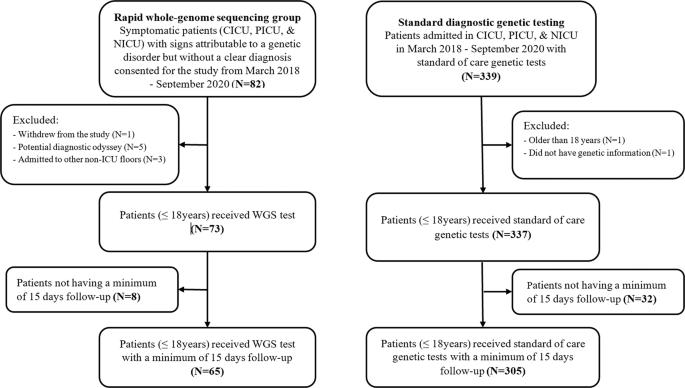
Real-world economic evaluation of prospective rapid whole-genome sequencing compared to a matched retrospective cohort of critically ill pediatric patients in the United States
There is an increasing demand for supporting the adoption of rapid whole-genome sequencing (rWGS) by demonstrating its real-world value. We aimed to assess the cost-effectiveness of rWGS in critically ill pediatric patients with diseases of unknown cause. Data were collected prospectively of patients admitted to the Nicklaus Children’s Hospital’s intensive care units from March 2018 to September 2020, with rWGS (N = 65). Comparative data were collected in a matched retrospective cohort with standard diagnostic genetic testing. We determined total costs, diagnostic yield (DY), and incremental cost-effectiveness ratio (ICER) adjusted for selection bias and right censoring. Sensitivity analyses explored the robustness of ICER through bootstrapping. rWGS resulted in a diagnosis in 39.8% while standard testing in 13.5% (p = 0.026). rWGS resulted in a mean saving per person of $100,440 (SE = 26,497, p < 0.001) and a total of $6.53 M for 65 patients. rWGS in critically ill pediatric patients is cost-effective, cost-saving, shortens diagnostic odyssey, and triples the DY of traditional approaches.
求助全文
通过发布文献求助,成功后即可免费获取论文全文。
去求助
来源期刊

Pharmacogenomics Journal
医学-药学
CiteScore
7.20
自引率
0.00%
发文量
35
审稿时长
6-12 weeks
期刊介绍:
The Pharmacogenomics Journal is a print and electronic journal, which is dedicated to the rapid publication of original research on pharmacogenomics and its clinical applications.
Key areas of coverage include:
Personalized medicine
Effects of genetic variability on drug toxicity and efficacy
Identification and functional characterization of polymorphisms relevant to drug action
Pharmacodynamic and pharmacokinetic variations and drug efficacy
Integration of new developments in the genome project and proteomics into clinical medicine, pharmacology, and therapeutics
Clinical applications of genomic science
Identification of novel genomic targets for drug development
Potential benefits of pharmacogenomics.
 求助内容:
求助内容: 应助结果提醒方式:
应助结果提醒方式:


